利用深度学习扩展双光子成像视场
Two-photon microscopy (TPM) imaging has been widely used in many fields, such as in vivo tumor imaging, neuroimaging, and brain disease research. However, the small field-of-view (FOV) in two-photon imaging (typically within diameter of 1 mm) limits its further application. Although the FOV can be extended through adaptive optics technology, the complex optical paths, additional device costs, and cumbersome operating procedures limit its promotion. In this study, we propose the use of deep learning technology instead of adaptive optics technology to expand the FOV of two-photon imaging. The large FOV of TPM can be realized without additional hardware (such as a special objective lens or phase compensation device). In addition, a BN-free attention activation residual U-Net (nBRAnet) network framework is designed for this imaging method, which can efficiently correct aberrations without requiring wavefront detection.
Commercially available objectives have a nominal imaging FOV that has been calibrated by the manufacturer. Within the nominal FOV, the objective lens exhibits negligible aberrations. However, the aberrations increase dramatically beyond the nominal FOV. Therefore, the imaging FOV of the objective lens is limited to its nominal FOV. In this study, we improved the imaging quality of the FOV outside the nominal region by combining adaptive optics (AO) and deep learning. Aberrant and AO-corrected images were collected outside the nominal FOV. Thus, we obtained a paired dataset consisting of AO-corrected and uncorrected images. A supervised neural network was trained using the aberrated images as the input and the AO-corrected images as the output. After training, the images collected from regions outside the nominal FOV could be fed directly to the network. Aberration-corrected images were produced, and the imaging system could be used without AO hardware.
The experimental test results include the imaging results of samples such as fluorescent beads with diameter of 1 μm and Thy1-GFP and CX3CR1-GFP mouse brain slices, and the results of the corresponding network output. The high peak signal-to-noise ratio (PSNR) values of the test output and ground truth demonstrate the feasibility of extending the FOV to TPM imaging using deep learning. At the same time, the intensity contrast between the nBRAnet network output image and the ground truth on the horizontal line is compared in detail (Figs. 3, 4, and 5). The extended FOV of different samples is randomly selected for analysis, and a high degree of coincidence is observed in the intensity comparison. The experimental results show that after using the network, both the resolution and fluorescence intensity can be restored to a level where there is almost no aberration, which is close to the result after correcting using AO hardware. To demonstrate the advantages of the network framework designed in this study, the traditional U-Net structure and the very deep super-resolution (VDSR) model are used to compare with ours. When using the same training dataset to train different models, we find that the experimental results of the VDSR model contain a considerable amount of noise, whereas the experimental results of the U-Net network lose some details (Fig. 6). The high PSNR values clearly demonstrate the strength of our nBRAnet network framework (Table 3).
This study provides a novel method to effectively extend the FOV of TPM imaging by designing an nBRAnet network framework. In other words, deep learning was used to enhance the acquired image and expand the nominal FOV for commercial objectives. The experimental results show that images from extended FOVs can be restored to their AO-corrected versions using the trained network. That is, deep learning technology could be used instead of AO hardware technology to expand the FOV of commercially available objectives. This simplifies the operation and reduces the system cost. An extended FOV obtained using deep learning can be employed for cross-regional or whole-brain imaging.
1 引言
双光子显微镜(TPM)具有成像分辨率高、深度大、三维层析能力强等成像特性,可为生物医学研究提供关键的高分辨三维信息,已被广泛应用于生物医学成像领域。然而,传统的双光子显微镜成像视场(FOV)有限,视场直径通常在1 mm以内[1-3],限制了其在更广泛的生物医学研究中的应用,如,双光子显微镜不能动态监测哺乳动物大脑中的跨区域神经元活动和全脑的血细胞活动[4-6]。因此,拓展双光子显微镜的成像视场一直是显微成像领域研究的难点和热点。
针对上述问题,研究人员尝试采用多种方法来提高双光子显微镜的成像视场,如:通过设计特殊的扫描中继系统[7]来减小大角度扫描引起的离轴像差,从而极大地优化大视场双光子成像系统的成像性能。此外,成像物镜也是限制视场进一步增大[8-10]的关键因素,通过自制高通量物镜[11-12]可以实现更大的成像视场。以上方法都可以有效地扩展成像视场,但较高实现难度、较大成本的光学设计严重制约了这些方法在生物医学研究领域的推广。近期,笔者团队[13]提出了一种通过自适应光学(AO)技术[14-18]增加成像物镜可用视场的新方法,该方法基于现有商业显微物镜在超过标定视场区域有信号但光学畸变严重这一特性,利用自适应光学技术分块补偿光学畸变,将商业物镜(XLPLN10XSVMP,Olympus,10×,NA=0.6)的有效成像视场直径从标定的1.8 mm扩展至3.46 mm,同时在全视场内保持了高分辨率和高信噪比成像。但是该方法需要在光学系统中增加相位补偿器件(空间光调制器),这无疑增加了成像系统的光路复杂性与硬件成本,从而限制了该方法的进一步应用。
利用计算机进行图像分辨率和对比度的提升是一种有效的降低硬件成本的方法。近年来,深度学习技术在图像处理方面发展迅速,已在图像恢复、超分辨率成像[19-23]、荧光显微图像重建[24-31]等领域得到了广泛应用。2021年,Zheng等[32]建立了前15项Zernike模式的系数与其对应的畸变模式之间的映射关系,通过卷积神经网络对光学像差引起的成像畸变进行了校正。2020年,Krishnan等[33]提出了一种深度学习方法,该方法通过联合Zernike系数估计、点扩散函数估计和盲解卷积训练的卷积神经网络(CNN)从3个相位差异图像中恢复无像差图像。2021年,Hu课题组[34]采用一种结合像差先验知识的尺度循环像差校正网络(SRACNet)来增强荧光显微镜图像,并提出了一种数据集生成策略,改善了数据集问题,实现了无须使用额外硬件的像差校正。以上研究主要利用深度学习结合自适应光学来提升成像分辨率,极大地降低了自适应光学技术的难度和成本。目前,利用深度学习技术代替自适应光学技术来扩展双光子成像视场的相关研究还未见公开报道。
本文提出了一种利用深度学习替代自适应光学相位补偿从而扩展双光子成像视场的新方法,该方法是对本团队近期发展的基于自适应光学技术扩展物镜有效视场方法的一次新改进[13]。该方法无须使用复杂的自适应光学元件进行像差校正,而且利用深度学习替代本团队近期所提方法中用硬件实现的自适应光学补偿,不仅降低了系统成本,实现了无须硬件自适应光学补偿的大视场成像,还提高了系统的便捷性。在本实验中,笔者设计了一种nBRAnet网络框架,该框架可以高效、便捷地增强双光子扩展视场的分辨率。采用直径为1 µm的荧光小球、Thy1-GFP小鼠脑切片、CX3CR1-GFP小鼠脑切片等样品的成像结果对所提网络进行训练和测试。实验结果表明:所提网络可以快速输出扩展视场的高分辨率图像,无论是分辨率还是荧光强度,都能恢复到近似无像差时的水平,即接近硬件自适应光学校正之后的结果。
2 方法与原理
2.1 拓展成像视场的原理
通常,商用物镜都有一个厂商标定的成像视场[如
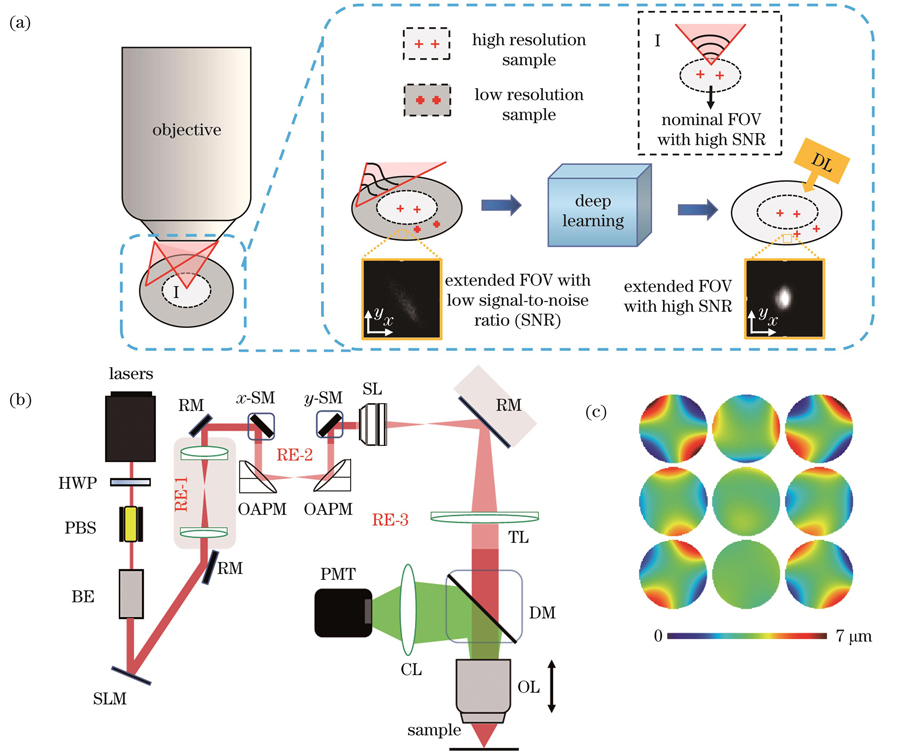
图 1. 光学系统及成像原理。(a)深度学习扩展双光子显微镜可用视场的原理图;(b)大视场双光子显微镜系统示意图(HWP:半波片;PBS:偏振分光棱镜;PMT:光电倍增管;RM:反射镜;BE:扩束器;SLM:空间光调制器;OAPM:离轴抛物面反射镜;SM:扫描振镜;SL:扫描透镜;TL:管透镜;DM:二向色镜;OL:物镜;CL:收集透镜);(c)测量得到的3×3个子区域的波前面
Fig. 1. Optical systems and imaging principle. (a) Principle of extending field-of-view (FOV) of two-photon microscopy (TPM) using deep learning; (b) schematic of TPM system with large FOV (HWP: half wave plate; PBS: polarization splitting prism; PMT: photomultiplier; RM: reflective mirror; BE: beam expander; SLM: spatial light modulator; OAPM: off-axis parabolic mirror; SM: scan mirror; SL: scan lens; TL: tube lens; DM: dichroic mirror; OL: objective lens; CL: collective lens); (c) measured distort wavefronts of 3×3 subregions
扩展成像视场的原理如
2.2 光学系统介绍与像差获取
本文使用的大视场双光子显微系统如
采用间接波前检测中的模式法[15]实现像差测量和校正。模式法的原理是:将波前畸变表示为一系列正交多项式之和,对每种模式像差进行单独校正。该方法常用Zernike多项式来表示基本像差模式,Zernike多项式是定义在单位圆上且满足正交的多项式序列[33]。
表 1. 本文所用各项Zernike多项式
Table 1. Applied Zernike polynomials
|
考虑到整个大视场中像差分布不均匀,因此将整个视场区域分为3×3个子区域,进行分区校正。对于每一个子区域,测量前已调整物镜的校正环,以最小化系统引入的球差。第1个Zernike模式(Piston)表示穿过光瞳的波前的平均值,对光斑没有影响;第2个和第3个Zernike模式(Tip和Til)引起了二维焦平面上的空间运动,但不影响分辨率或信号强度。所以,本文使用Zernike多项式的第5~15项进行像差计算。每个分区的各项最优系数的测量方法一致:从第5项开始依次测量每一项的最优系数,测量过程中将前一项的最优系数作为下一项系数测量的基准。考虑到扩展视场有较大的离轴像差,因此先单独确定每块区域中的像散对应的Zernike系数,即:将系数从-20改变为20,间隔为0.5,使用SLM连续加载对应的波前,并对激发荧光均匀片产生的双光子图像进行采集,对每幅图的平均光强进行拟合和搜索,得到最大值对应的系数值即为所求的像散系数。对于该区域其他被应用的Zernike多项式,将系数从-3改变为3,间隔为0.5,同样通过采集双光子图像得到最佳系数。最终所有阶数的Zernike像差模式的线性叠加即为相应分区的整体像差。以上步骤重复测量三次后取平均,以保证测量准确度。每块分区对应的波前面如
2.3 数据准备
数据支撑[27]是深度学习中图像复原任务的最核心问题之一。数据集的匹配程度决定了神经网络的可靠性,为此本文作出了两方面改进:1)提高信噪比。由于单次测量信噪比不足,所以对每组校正前后的图像各采集3张,并通过平均图像的灰度值来提高图像的信噪比。2)匹配图像的三维位置。原始图像在自适应光学校正前后会有轻微的像素不匹配,因此,本文将自适应光学技术校正前的图像作为参照,通过配准算法对自适应光学校正后的图像进行X和Y方向的配准,将配准后的有效数据尺寸裁剪为1000 pixel×1000 pixel。
本文中的训练数据集均由
2.4 nBRAnet网络结构
本实验采用的基于U-Net改进的nBRAnet网络如

图 2. 本文提出的nBRAnet网络结构。(a)改进的3层网络结构;(b)残差结构示意图;(c)上采样结构块;(d)改进的卷积块;
Fig. 2. Proposed nBRAnet network structure. (a) Improved three-layer network structure; (b) schematic of residual structure;
考虑到网络深度会导致训练困难的问题,本工作在U-Net网络框架[35]的基础上引入残差结构[36][如
表 2. 验证BN结构影响的消融实验的结果
Table 2. Results from ablation experiments for verifying BN structure’s effect
|
2.5 网络训练设置
神经网络的训练在Windows环境下进行,采用PyTorch(Python3.7)编写代码,使用MATLAB代码处理图片。实验在桌面工作站(Intel Xeon E5-2620 v4 CPU,NVIDIA TITAN Xp)上进行。数据集由荧光小球和离体生物样品经自适应光学校正前后的显微图像构成,网络模型输入图像和输出图像的尺寸均为1000 pixel×1000 pixel。分别取数据集的95%和5%用作训练集和测试集。利用网络输出与标签之间的均方误差(MSE)作为损失函数来训练模型,同时利用反向传播算法Adam来优化网络,初始学习速率为1×10-4。网络模型的参数总量为3.558×107。
3 结果与讨论
3.1 荧光小球实验结果
为了定量评估网络的输出结果,使用峰值信噪比(PSNR)作为评价指标。PSNR是一种衡量图像质量的指标,常被用作图像降噪领域信号重建质量的重要指标,其值越高说明重建结果越好。
首先,使用荧光小球仿体样品验证深度学习方法可用于扩展双光子成像视场。

图 3. 直径为1 µm的荧光小球的大视场成像结果。(a)全视场荧光小球图像,视场尺寸为2.45 mm×2.45 mm;(b)~(d)标定区域经自适应光学校正前后的图像以及网络模型学习得到的图像;(e)~(g)扩展区域经自适应光学校正前后的图像以及网络模型学习得到的图像;(h)Ⅰ区域的强度曲线;(i)Ⅱ区域的强度曲线
Fig. 3. Large FOV imaging of fluorescent beads with diameter of 1 µm. (a) Image of fluorescent beads with the FOV of 2.45 mm×2.45 mm; (b)-(d) images with and without adaptive optics (AO) correction as well as obtained by network model learning in nominal FOV; (e)-(g) images with and without AO correction as well as obtained by network model learning in extended FOV; (h) intensity profiles of region Ⅰ; (i) intensity profiles of region Ⅱ
3.2 生物样品实验结果
为了进一步评估深度学习对双光子显微镜视场扩展的可行性,将本文所提网络应用于不同离体生物样品的成像实验中。脑科学研究是大视场双光子显微镜应用的重要领域,所以优先选择Thy1-GFP小鼠脑片进行成像分析。

图 4. Thy1-GFP小鼠大脑切片的大视场成像结果。(a)全视场图像,视场尺寸为2.45 mm×2.45 mm;(b)虚线框区域扩展视场在自适应光学校正前的图像;(c)虚线框区域扩展视场在自适应光学校正后的图像;(d)深度学习模型的图像增强结果;(e)(f)划线区域的强度对比
Fig. 4. Large FOV imaging of brain slice of Thy1-GFP mouse. (a) Full FOV image with size of 2.45 mm×2.45 mm; (b) image of extended FOV in dash box before AO correction; (c) image of extended FOV in dash box after AO correction; (d) enhanced image obtained by deep learning model; (e)(f) intensity comparison along solid and dash lines
为验证网络的通用性,使用GFP荧光标记的小胶质细胞进行进一步验证(Thy1-GFP主要标记的是神经元细胞,CX3CR1-GFP主要标记的是小胶质细胞,两者形态有较大差异)。

图 5. CX3CR1-GFP小鼠大脑切片中小胶质细胞的大视场成像。(a)全视场图像,视场尺寸为2.45 mm×2.45 mm;(b)虚线框区域扩展视场在自适应光学校正前的图像;(c)虚线框区域扩展视场在自适应光学校正后的图像;(d)所提深度学习模型的图像增强结果;(e)~(g)灰度值直方图,分别对应(b)~(d)
Fig. 5. Large FOV imaging of microglia in CX3CR1-GFP mouse brain slice. (a) Full FOV image with size of 2.45 mm×2.45 mm; (b) image of extended FOV in dash box before AO correction; (c) image of extended FOV in dash box after AO correction; (d) enhanced image obtained by deep learning model; (e)-(g) histograms of gray values corresponding to Figs.(b)-(d), respectively
不同的生物样品实验结果表明,深度学习技术不仅可以有效地替代硬件自适应光学技术,增强双光子显微镜扩展视场中生物结构的特征,还可以大幅降低图像中的噪声,恢复成像分辨率。
3.3 网络评估
为展示本文改进网络框架的优势,将其与超深超分辨率模型(VDSR)[40]、传统的U-Net模型进行比较。nBRAnet、VDSR和U-Net分别采用相同的数据集进行训练,不同网络模型的输出结果如

图 6. 不同网络模型的输出结果。(a)荧光小球样品扩展视场的ROI区域;(b)Thy1-GFP样品扩展视场的ROI区域;(c)CX3CR1-GFP样品扩展视场的ROI区域
Fig. 6. Output results of different network models. (a) ROI area of extended FOV of fluorescent bead samples; (b) ROI area of extended FOV of Thy1-GFP sample; (c) ROI area of extended FOV of CX3CR1-GFP sample
表 3. 不同网络模型的实验结果评估
Table 3. Experimental results evaluation of different network models
| ||||||||||||||||||||||||||||||||||||||||||||||
需要指出的是,在这项工作中,改进的模型在U-Net框架的基础上添加了残差结构和空间注意力结构,使得网络框架更加复杂,而且该模型去除了BN结构,减缓了网络的收敛速度,导致改进后的模型参数总量增加,在提升精度的同时牺牲了一定的时间。另外,训练数据集分别为自适应光学校正前后的二维图像,但在实际采样过程中,由于校正前的点扩散函数沿z轴方向的长度更长(相比于校正后),自适应光学校正前的图像叠加了其他z轴位置的信息,使得校正前后的图像不完全匹配。为解决这一问题,后续拟考虑使用三维数据代替二维数据。三维数据的优势在于其通过z轴多层采样后,网络的输入输出包含了更完整的样品特征,从而更好地还原了样品信息。此外,深度学习的泛化能力还有待于进一步加强。在实验过程中,笔者曾尝试将小球的训练模型应用在其他生物样品上,但网络输出的效果并不理想。这也是深度学习依赖数据驱动的局限性。这就要求采集的数据集除了清晰、特征明显外,还需要包含更多类型的样品图像,使其种类和特征能够覆盖将来潜在的测试样品,以此提高网络模型的性能和泛化能力。
此外,笔者除了在文内展示的Thy1-GFP大脑切片样品和CX3CR1-GFP大脑切片样品上进行了成像研究之外,还对多个样品进行了实验。不同样品经网络恢复后,其扩展视场区域的分辨率和荧光强度均得到较大提升,均能近似地恢复到硬件自适应光学校正后无像差时的水平。例如:植物细胞样品扩展视场的ROI区域经网络恢复后,图像的信噪比由原来的15.75 dB提升至20.83 dB;不同小鼠的Thy1-GFP大脑切片样品扩展视场的ROI区域经网络恢复后,图像的信噪比由原来的30.58 dB提升至35.45 dB。今后将继续尝试更多的生物样品。
4 结论
本文提供了一种有效扩展双光子显微镜成像视场的新思路、新途径,即,利用深度学习来扩展商业物镜的可用视场。通过分析荧光小球实验、Thy1-GFP小鼠脑内神经细胞实验和CX3CR1-GFP小鼠脑内小胶质细胞实验可知,经网络恢复后的扩展区域图像,无论是分辨率还是荧光强度,均能恢复到接近硬件自适应光学校正后无像差时的水平。换言之,本文所提方法利用训练好的模型可以直接输出校正像差后的样品扩展视场的ROI区域图像,无须用硬件自适应元件进行像差校正,简化了操作,降低了系统的复杂度和成本,提高了成像分辨率及其拓展的通用性,具有较高的实用价值。
[1] Zipfel W R, Williams R M, Webb W W. Nonlinear magic: multiphoton microscopy in the biosciences[J]. Nature Biotechnology, 2003, 21(11): 1369-1377.
[2] König K. Multiphoton microscopy in life sciences[J]. Journal of Microscopy, 2000, 200(2): 83-104.
[3] Helmchen F, Denk W. Deep tissue two-photon microscopy[J]. Nature Methods, 2005, 2(12): 932-940.
[4] Bumstead J R, Park J J, Rosen I A, et al. Designing a large field-of-view two-photon microscope using optical invariant analysis[J]. Neurophotonics, 2018, 5(2): 025001.
[5] Ji N, Freeman J, Smith S L. Technologies for imaging neural activity in large volumes[J]. Nature Neuroscience, 2016, 19(9): 1154-1164.
[6] Yang W J, Yuste R. In vivo imaging of neural activity[J]. Nature Methods, 2017, 14(4): 349-359.
[7] Tsai P S, Mateo C, Field J J, et al. Ultra-large field-of-view two-photon microscopy[J]. Optics Express, 2015, 23(11): 13833-13847.
[8] Yu C H, Stirman J N, Yu Y Y, et al. Diesel2p mesoscope with dual independent scan engines for flexible capture of dynamics in distributed neural circuitry[J]. Nature Communications, 2021, 12(1): 1-8.
[9] Clough M, Chen I A, Park S W, et al. Flexible simultaneous mesoscale two-photon imaging of neural activity at high speeds[J]. Nature Communications, 2021, 12(1): 1-7.
[10] Demas J, Manley J, Tejera F, et al. High-speed, cortex-wide volumetric recording of neuroactivity at cellular resolution using light beads microscopy[J]. Nature Methods, 2021, 18(9): 1103-1111.
[11] Sofroniew N J, Flickinger D, King J, et al. A large field of view two-photon mesoscope with subcellular resolution for in vivo imaging[J]. eLife, 2016, 5: e14472.
[12] Stirman J N, Smith I T, Kudenov M W, et al. Wide field-of-view, multi-region two-photon imaging of neuronal activity in the mammalian brain[J]. Nature Biotechnology, 2016, 34(8): 857-62.
[13] Yao J, Gao Y F, Yin Y X, et al. Exploiting the potential of commercial objectives to extend the field of view of two-photon microscopy by adaptive optics[J]. Optics Letters, 2022, 47(4): 989-992.
[14] 刘立新, 张美玲, 吴兆青, 等. 自适应光学在荧光显微镜中的应用[J]. 激光与光电子学进展, 2020, 57(12): 120001.
[15] Booth M J, Neil M A A, Juskaitis R, et al. Adaptive aberration correction in a confocal microscope[J]. Proceedings of the National Academy of Sciences of the United States of America, 2002, 99(9): 5788-5792.
[16] Park J H, Kong L J, Zhou Y F, et al. Large-field-of-view imaging by multi-pupil adaptive optics[J]. Nature Methods, 2017, 14(6): 581-583.
[17] Ji N. Adaptive optical fluorescence microscopy[J]. Nature Methods, 2017, 14(4): 374-380.
[18] Booth M J. Adaptive optical microscopy: the ongoing quest for a perfect image[J]. Light: Science & Applications, 2014, 3(4): e165.
[19] Zhou H, Cai R Y, Quan T W, et al. 3D high resolution generative deep-learning network for fluorescence microscopy imaging[J]. Optics Letters, 2020, 45(7): 1695-1698.
[20] Zhang H, Fang C Y, Xie X L, et al. High-throughput, high-resolution deep learning microscopy based on registration-free generative adversarial network[J]. Biomedical Optics Express, 2019, 10(3): 1044-1063.
[21] Weigert M, Schmidt U, Boothe T, et al. Content-aware image restoration: pushing the limits of fluorescence microscopy[J]. Nature Methods, 2018, 15(12): 1090-1097.
[22] Cheng S F, Zhou Y Y, Chen J B, et al. High-resolution photoacoustic microscopy with deep penetration through learning[J]. Photoacoustics, 2022, 25: 100314.
[23] Zhao H X, Ke Z W, Chen N B, et al. A new deep learning method for image deblurring in optical microscopic systems[J]. Journal of Biophotonics, 2020, 13(3): e201960147.
[24] Belthangady C, Royer L A. Applications, promises, and pitfalls of deep learning for fluorescence image reconstruction[J]. Nature Methods, 2019, 16(12): 1215-1225.
[25] 李浩宇, 曲丽颖, 华子杰, 等. 基于深度学习的荧光显微成像技术及应用[J]. 激光与光电子学进展, 2021, 58(18): 1811007.
[26] Liu J H, Huang X S, Chen L Y, et al. Deep learning–enhanced fluorescence microscopy via degeneration decoupling[J]. Optics Express, 2020, 28(10): 14859-14873.
[27] 熊子涵, 宋良峰, 刘欣, 等. 基于深度学习的荧光显微性能提升(特邀)[J]. 红外与激光工程, 2022, 51(11): 89-106.
Xiong Z H, Song L F, Liu X, et al. Performance enhancement of fluorescence microscopy by using deep learning(invited)[J]. Infrared and Laser Engineering, 2022, 51(11): 89-106.
[28] Shen B L, Liu S W, Li Y P, et al. Deep learning autofluorescence-harmonic microscopy[J]. Light: Science & Applications, 2022, 11(1): 1-14.
[29] Wang Z Q, Zhu L X, Zhang H, et al. Real-time volumetric reconstruction of biological dynamics with light-field microscopy and deep learning[J]. Nature Methods, 2021, 18(5): 551-556.
[32] Zheng Y, Chen J J, Wu C X, et al. Adaptive optics for structured illumination microscopy based on deep learning[J]. Cytometry, 2021, 99(6): 622-631.
[34] Hu L J, Hu S W, Gong W, et al. Image enhancement for fluorescence microscopy based on deep learning with prior knowledge of aberration[J]. Optics Letters, 2021, 46(9): 2055-2058.
[35] RonnebergerO, FischerP, BroxT. U-Net: convolutional networks for biomedical image segmentation[M]∥Navab N, Hornegger J, Wells W M, et al. Medical image computing and computer-assisted intervention-MICCAI 2015. Lecture notes in computer science. Cham: Springer, 2015: 234-241.
[36] HeK M, ZhangX Y, RenS Q, et al. Deep residual learning for image recognition[C]∥2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), June 27-30, 2016, Las Vegas, NV, USA. New York: IEEE Press, 2016: 770-778.
[39] nBRAnet[EB/OL]. [2022-10-08]. https://gitee.com/li-chijian/nBRAnet.
[40] KimJ, LeeJ K, LeeK M. Accurate image super-resolution using very deep convolutional networks[C]∥2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), June 27-30, 2016, Las Vegas, NV, USA. New York: IEEE Press, 2016: 1646-1654.
Article Outline
李迟件, 姚靖, 高玉峰, 赖溥祥, 何悦之, 齐苏敏, 郑炜. 利用深度学习扩展双光子成像视场[J]. 中国激光, 2023, 50(9): 0907107. Chijian Li, Jing Yao, Yufeng Gao, Puxiang Lai, Yuezhi He, Sumin Qi, Wei Zheng. Extending Field‑of‑View of Two‑Photon Microscopy Using Deep Learning[J]. Chinese Journal of Lasers, 2023, 50(9): 0907107.