Photonics Research, 2019, 7 (9): 09001042, Published Online: Aug. 13, 2019
Simultaneous dual-contrast three-dimensional imaging in live cells via optical diffraction tomography and fluorescence  Download: 644次
Download: 644次
Figures & Tables
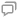
Fig. 1. Comparison of the 2D and 3D diffracted field using Fourier relationship. (a) 3D light field of a cell, shown as different planes of wavefront information; (b) 2D light field of a cell extracted from (a); (c) 3D and (d) 2D Fourier spectrum of the light fields. Under Fourier operation, a “slice” of the diffracted field results in a 2D spectrum as shown in (d), whereas the entire volumetric diffracted field results in a 3D “spherical cap,” as shown in (c).
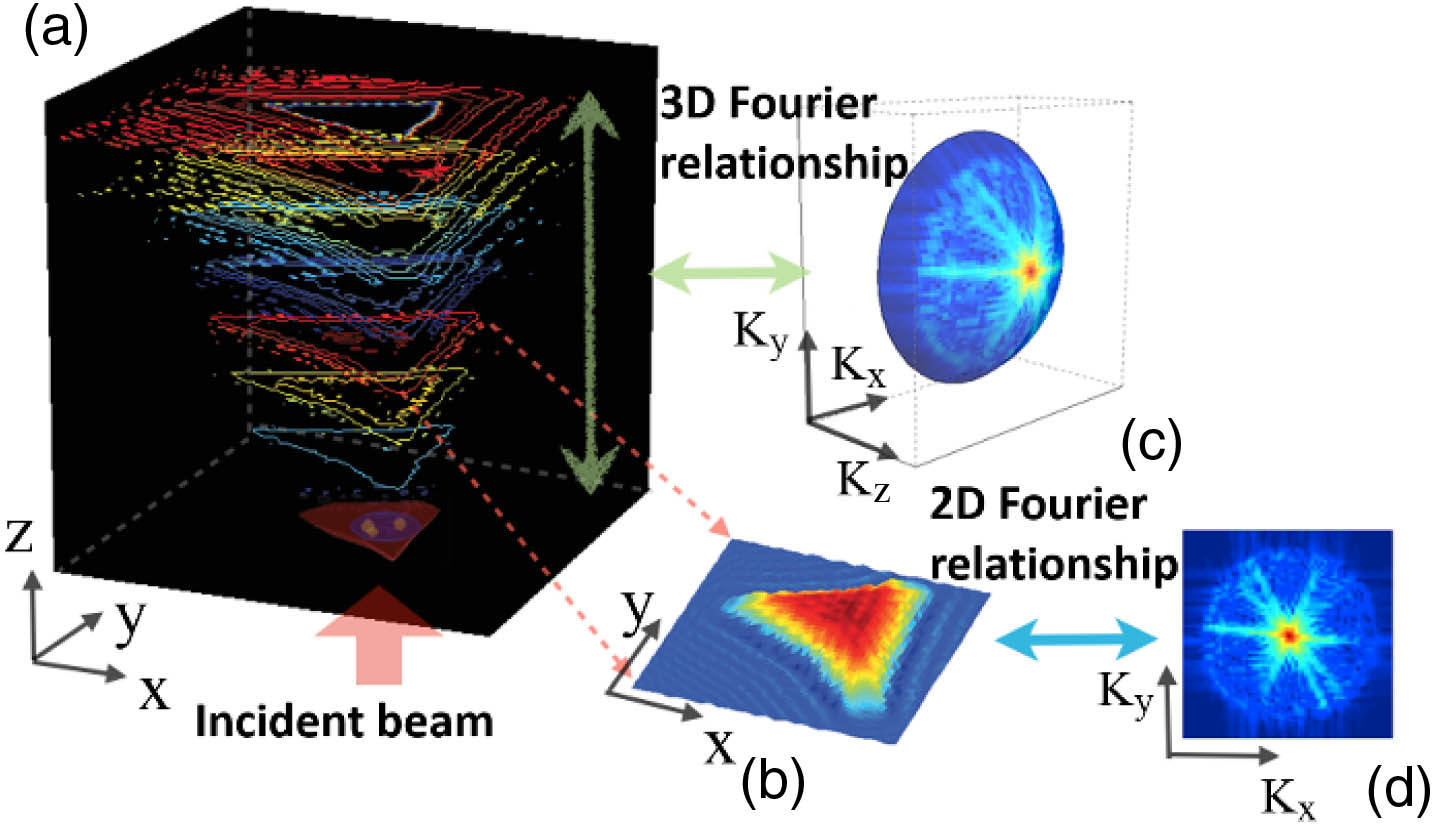
Fig. 2. Complex spectra synthesis of the scattering potential using three example angles. (a) A cell is subject to a multiangle plane-wave illumination with a maximum subtended half-angle θ N.A. = sin θ θ − θ Δ F z θ
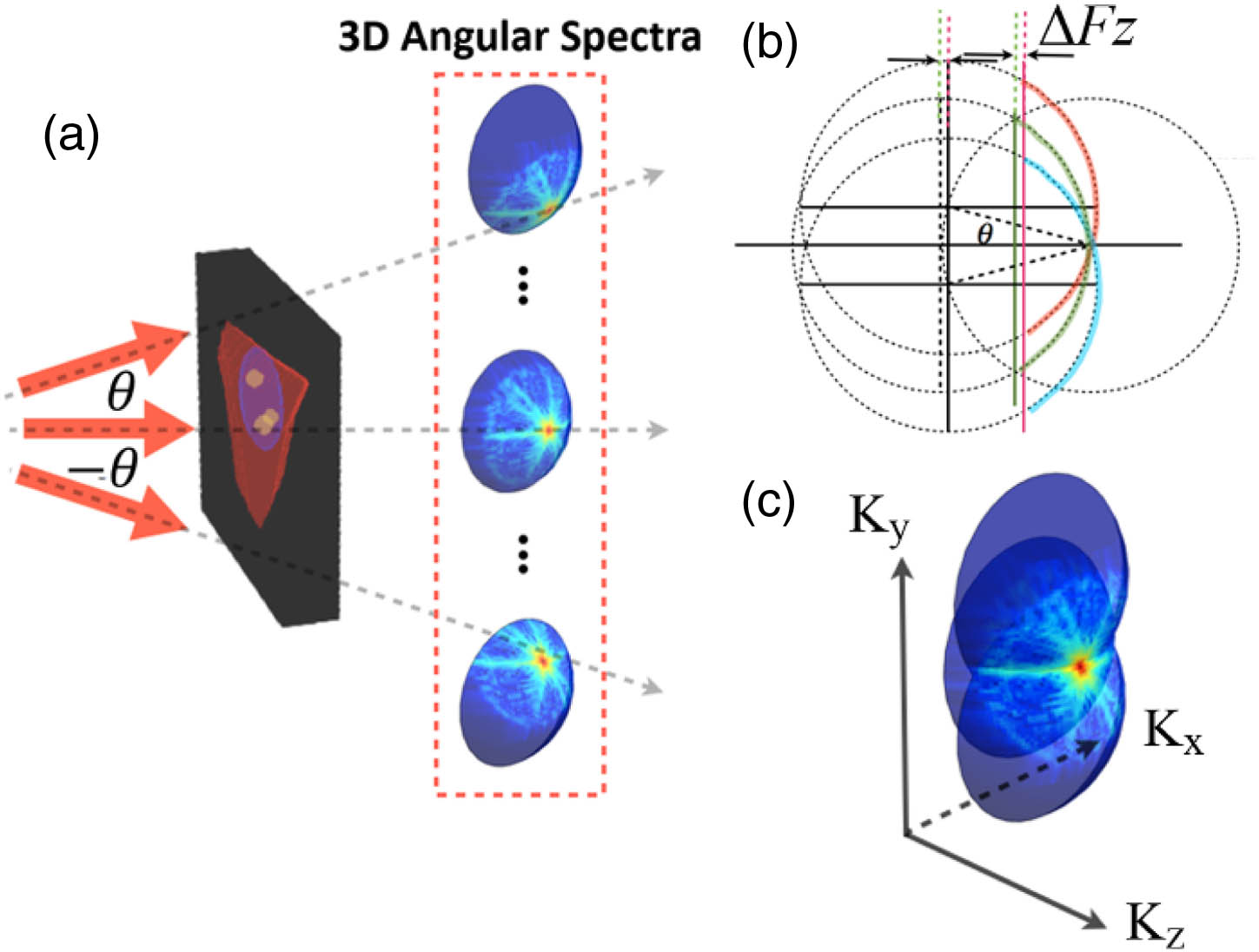
Fig. 3. Schematic diagram of the multimodal setup combining tomography with fluorescence. The tomography module is based on an off-axis digital holographic microscope setup in transmission. Different illumination angles are introduced by galvo mirror. The inset illustrates the tilted angle illumination situation. The fluorescence module is based on the epifluorescence microscopy. BS, beam splitter; GM, galvo mirror; SL, scanning lens; C, condenser; O, objective lens; TL, tube lens; Ex, excitation filter; Em, emission filter; DM, dichroic mirror.
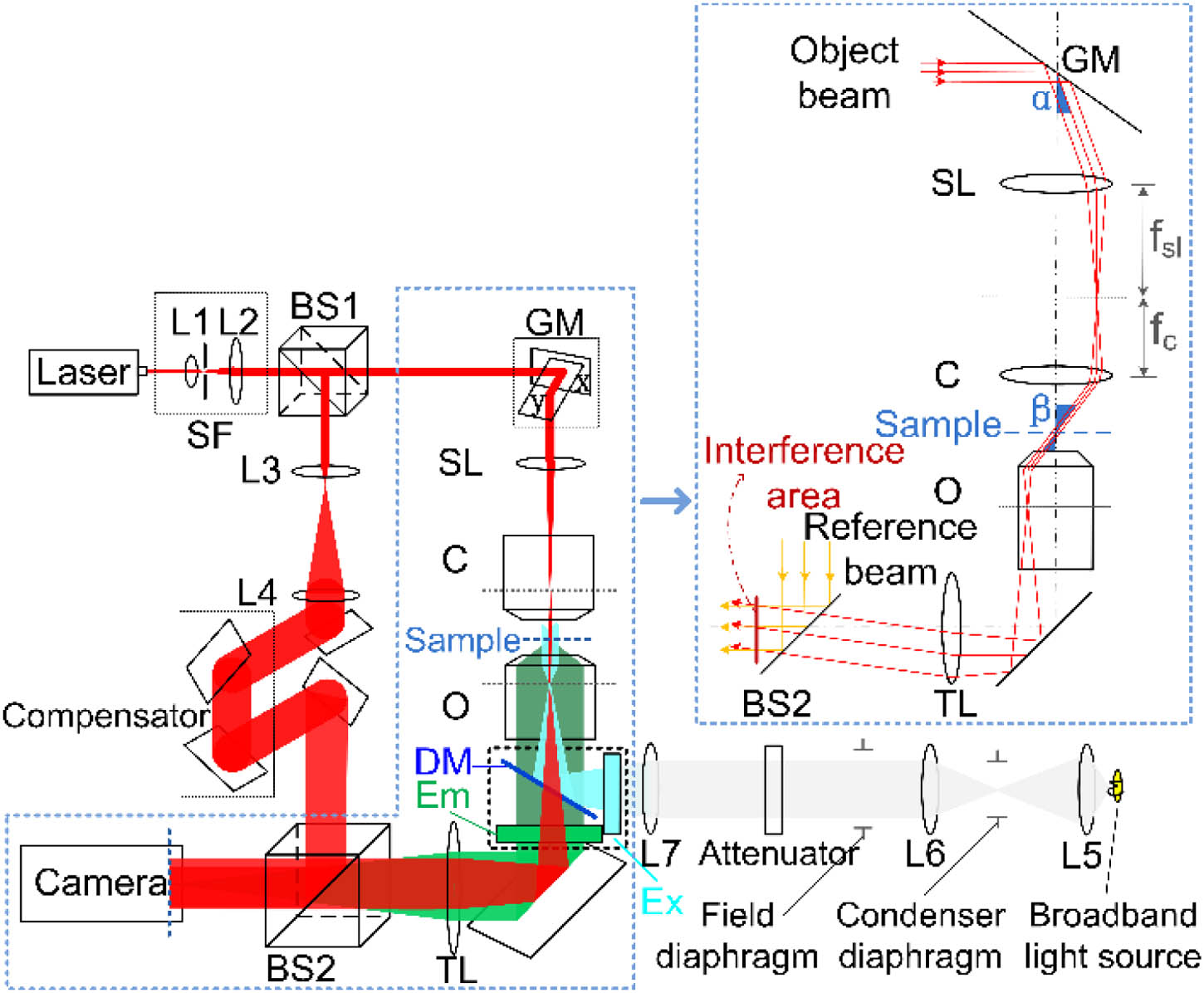
Fig. 4. Results for 3D reconstructions. (a) Buccal cell and (c) neuron, with internal structures delineated by groupings of refractive indices. The rotating views of the buccal cells and neuron can be found in Visualization 1 and Visualization 2 , respectively. Optical phase distribution of the same (b) buccal cell (in dashed box) and (d) neuron calculated from the interferometric measurements. The units for the color bars in (b) and (d) are radians. Part of the buccal cell in (a) is removed for clear visualization of the subcellular structures. The magnification is 20 ×
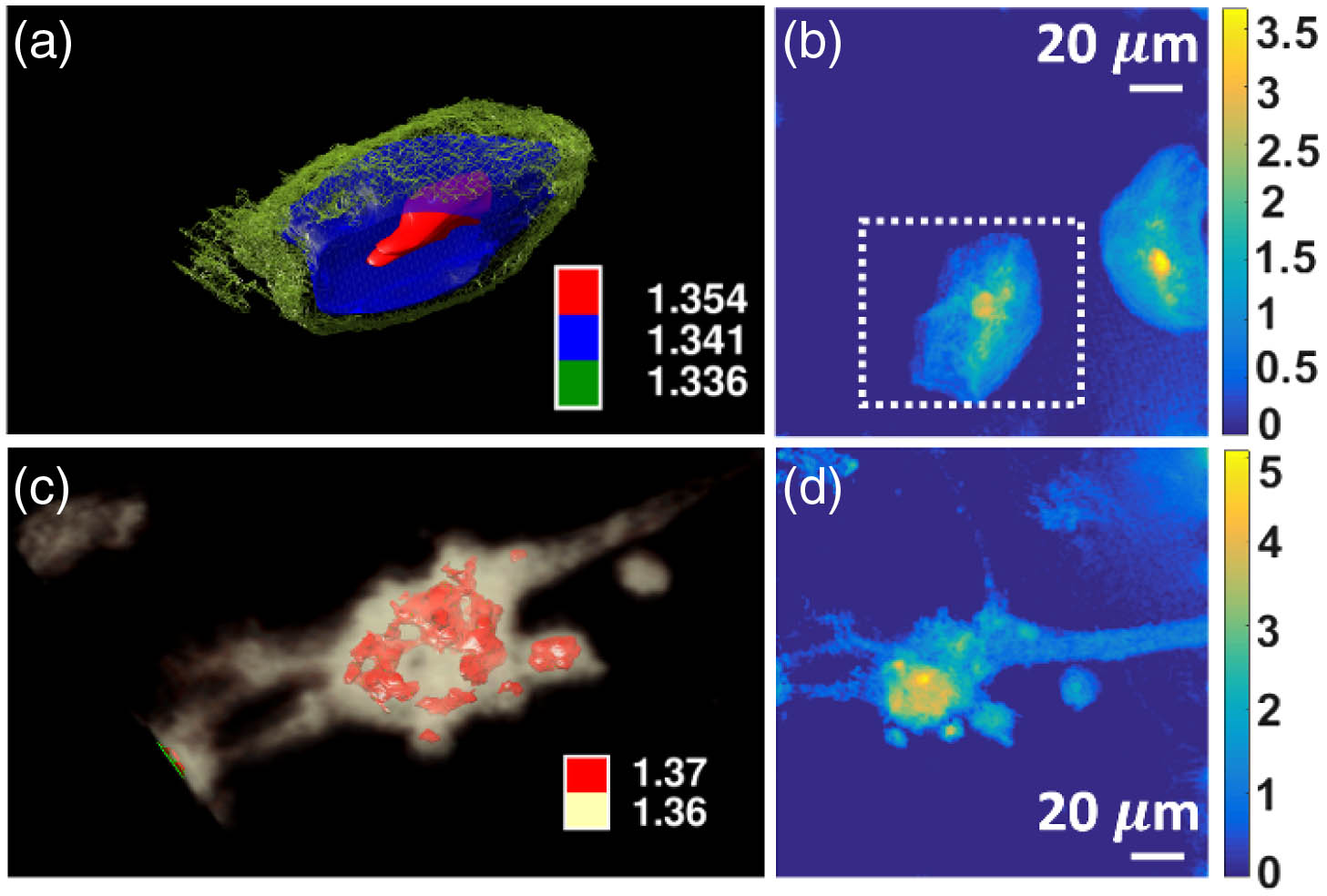
Fig. 5. Structures reconstructed using RI spatial mappings in apoptotic T cells are shown to be compatible with wide-field fluorescence counterparts. (a) Cells with internal structures delineated from groupings of RI values (top view). Inset gives an opened-up view of the cell through the central z Visualization 3 . (b) 2D slice of RI distributions along x y 40 ×
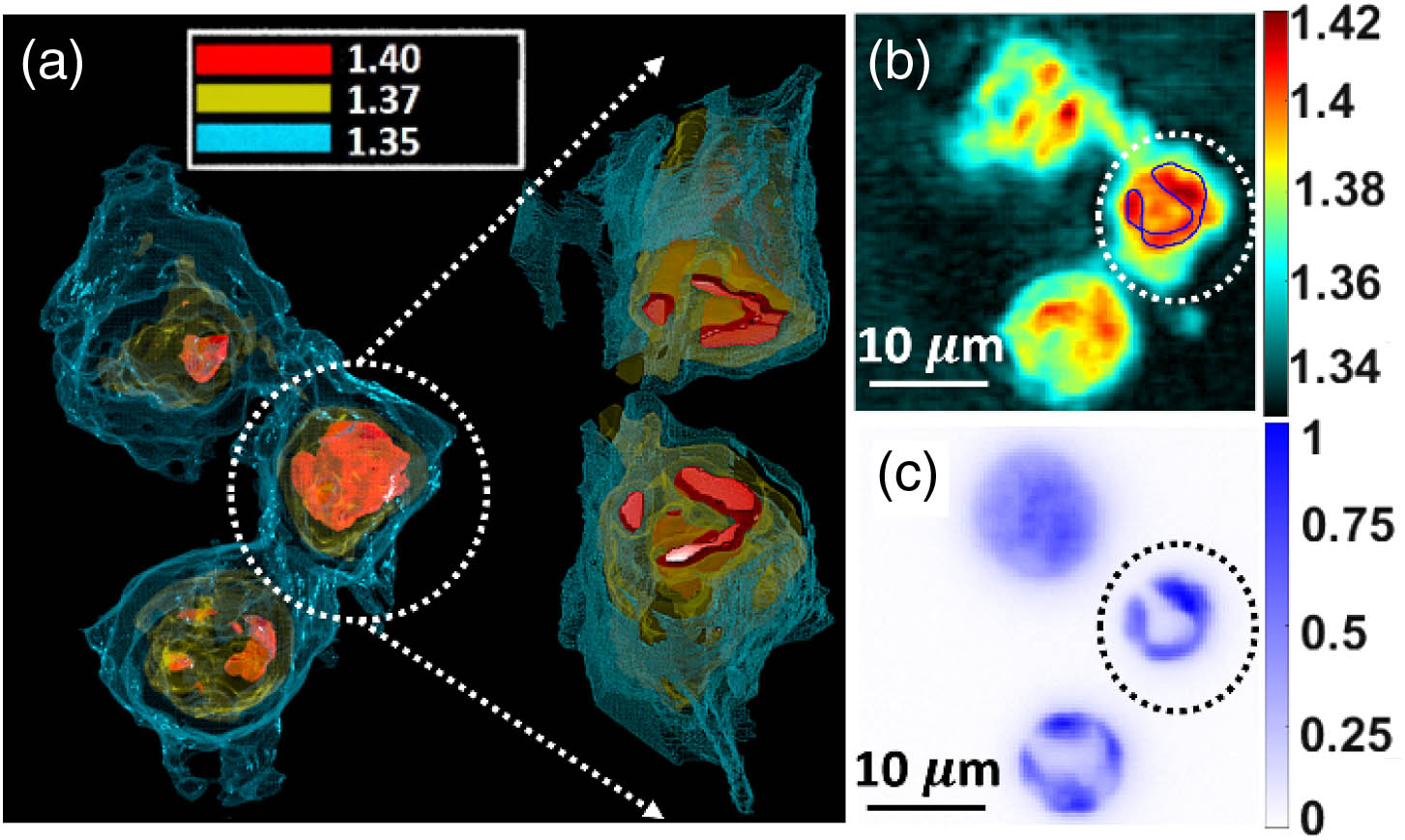
Fig. 6. Distributions of RI values (row 1) and fluorescence intensities (row 2) for Jurkat T cells under different conditions (scale bar, 10 μm). (a), (b) Viable cell; (c), (d) UV light-induced apoptotic cells with trovafloxacin treatment; (e), (f) necrotic cell. The positions of stained DNA, where many proteins are localized, are highlighted with blue contour lines in RI mappings based on fluorescence imaging. The magnification is 40 ×
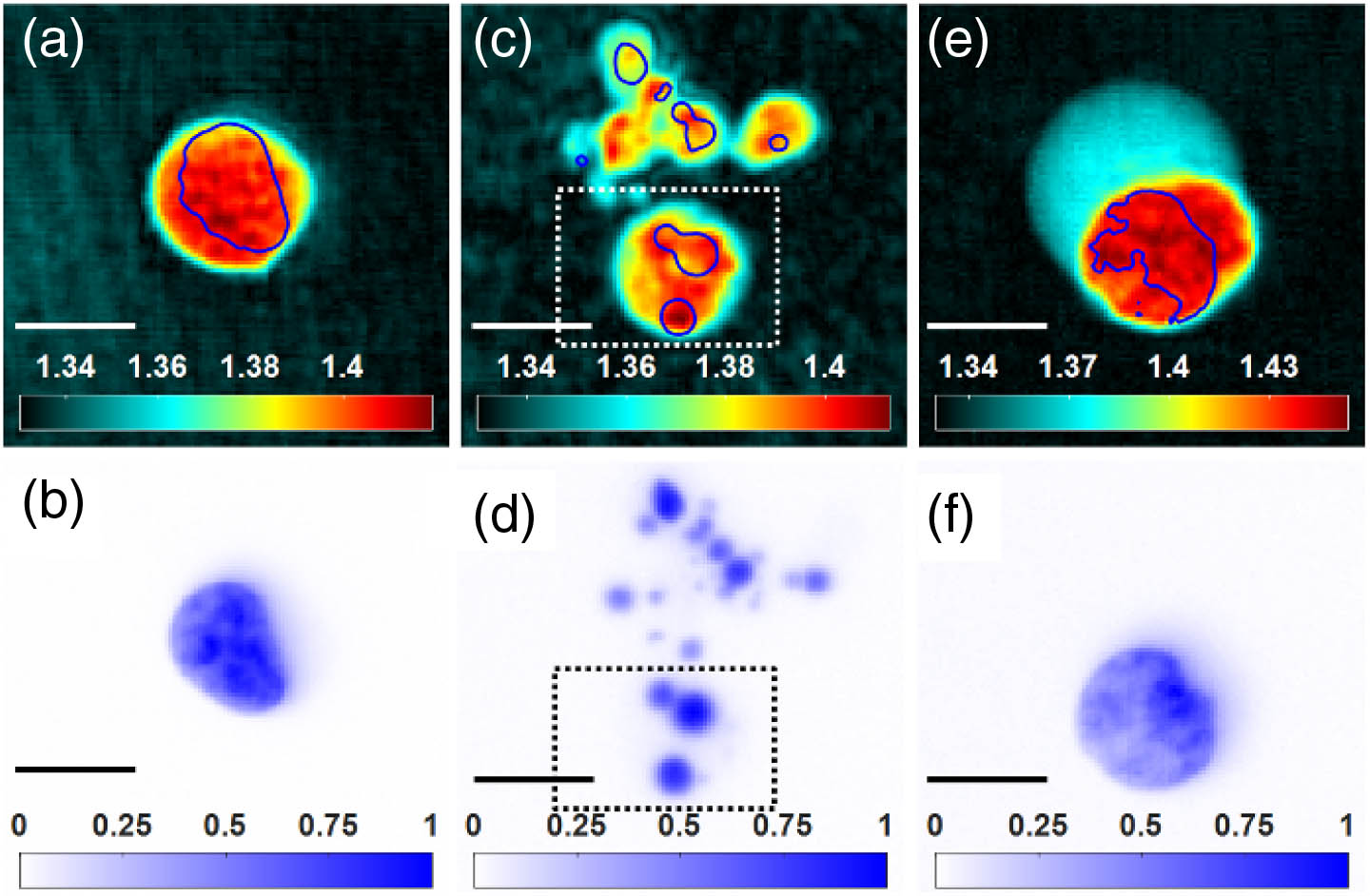
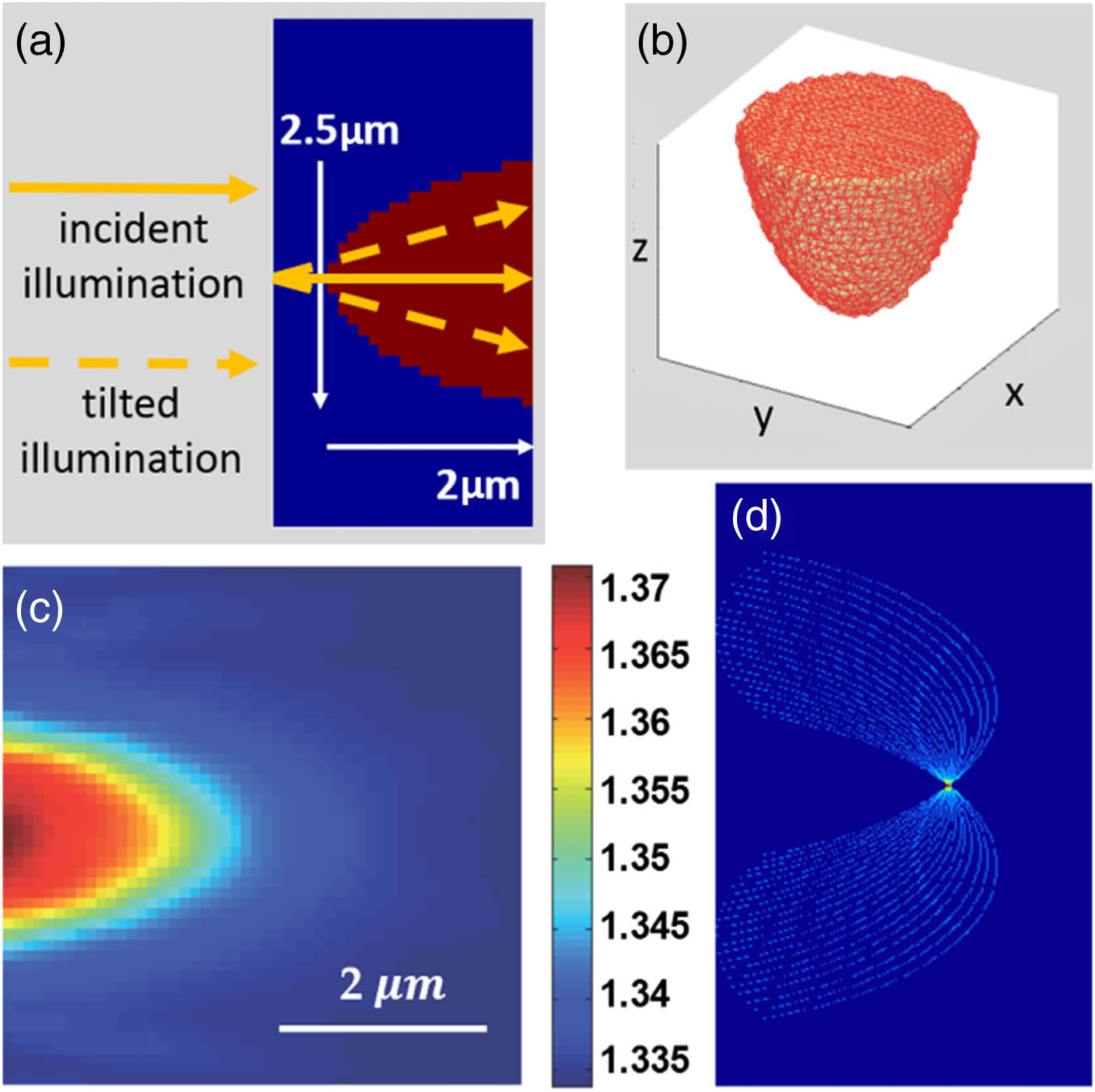
Fig. 7. Monte Carlo simulation of a phantom object volumetric diffraction and its quantitative reconstruction. (a) The configuration of the Monte Carlo simulation; (b) 3D view of the semispherical-shaped phantom object; (c) reconstructed RI distribution of the object in central y z f x f z
Chen Liu, Michael Malek, Ivan Poon, Lanzhou Jiang, Arif M. Siddiquee, Colin J. R. Sheppard, Ann Roberts, Harry Quiney, Douguo Zhang, Xiaocong Yuan, Jiao Lin, Christian Depeursinge, Pierre Marquet, Shan Shan Kou. Simultaneous dual-contrast three-dimensional imaging in live cells via optical diffraction tomography and fluorescence[J]. Photonics Research, 2019, 7(9): 09001042.





