基于Transformer的宫颈异常细胞自动识别方法
Cervical cancer is one of the most common malignant tumors and poses a serious threat to human health. However, because the onset of cervical cancer is gradual, early and effective screening is crucial. Traditional screening methods rely on manual examinations by pathologists, a process that is time-consuming, labor-intensive, error-prone, and often lacks an adequate number of pathologists for cervical cytology screening, making it challenging to meet the current demands for cervical cancer screening. In recent years, several deep-learning-based methods have been developed for screening abnormal cervical cells. However, because abnormal cervical cells develop from normal cells, they exhibit morphological similarities, making differentiation challenging. Pathologists typically need to reference normal cells in images to accurately distinguish them from abnormal cells. These factors limit the accuracy of abnormal cervical cell screening. This study proposes a Transformer-based approach for abnormal cervical cell screening that leverages the powerful global feature extraction and long-range dependency capabilities of Transformer. This method effectively enhances the detection accuracy of abnormal cervical cells, improving screening efficiency and alleviating the burden on medical professionals.
This study introduces a novel Transformer-based method for abnormal cervical cell detection that leverages the powerful global information extraction capabilities of Transformer to mimic the screening process of pathologists. The proposed method incorporates two innovative structures. The first is an improved Transformer encoder, which consists of multiple blocks stacked together. Each block comprises two parts: a multi-head self-attention layer and a feedforward neural network layer. The multi-head self-attention layer captures the correlation of the input data at different levels and scales, enabling the model to better understand the structure of the input sequence. The feedforward neural network layer includes multiple fully connected layers and activation functions and introduces nonlinear transformations to help the model adapt to complex data distributions. We also introduce Depthwise (DW)convolution and Dropout layers to the encoder. DW convolution layer performs convolution operations with separate kernels for each input channel, capturing features within the channels without introducing inter-channel dependencies. Dropout layer reduces the tendency of neural networks to overfit the training data, thereby enhancing the generalization of the model to unseen data. Additionally, we design a dynamic intersection-over-union (IOU) threshold method that adaptively adjusts the IOU threshold. In the initial stages of training, the model can obtain as many effective detections as possible, whereas in later stages, it can filter out most false positive predictions, thereby improving the detection accuracy of the model. Using the proposed method, the model can obtain precise information regarding the location of abnormal cells.
To validate the effectiveness of our proposed method, we compare it with common general-purpose object detection methods. The average accuracy (AP) and AP50 of our method are 26.1% and 46.8%, respectively, surpassing those of all general object detection models (Table 1). In particular, our method outperforms other comparative models by a significant margin in AP metrics, demonstrating that our model not only detects normal-sized targets but can also detect extremely small targets. Additionally, in a comparison with attFPN, a network specifically designed for abnormal cervical cell detection, our method surpasses attFPN in terms of AP by 1.1% (Table 2). Visual inspection of the detection results reveals that our method more accurately identifies target regions with lower false-positive and false-negative rates (Fig.5). Ablation experiments indicate that adopting the improved Transformer encoder method increases AP and AP50 by 1.8% and 2.3%, respectively, compared with the original model. The use of dynamic IOU thresholds results in a 0.6% increase in AP and a 0.9% increase in AP50 compared with the original model (Table 4). Furthermore, a comparison between the dynamic and fixed IOU thresholds in terms of loss and AP during the training process shows that the model with dynamic IOU thresholds experiences a faster loss reduction and achieves a higher AP in the later stages of training (Fig.6).
This study introduces an automatic identification method for abnormal cervical cells utilizing Transformer as the backbone. We further propose an enhanced Transformer encoder structure and a dynamically adjustable IOU threshold. Various comparative experiments on datasets demonstrate that the proposed method outperforms existing approaches in terms of accuracy and other metrics, thereby achieving precise identification of abnormal cervical cells. Through ablation experiments, it is proven that both proposed modules enhance the accuracy of the model in identifying abnormal cervical cells. Overall, the proposed method significantly improves the efficiency of medical image screening, saving medical time and resources, facilitating timely detection of cancerous lesions, and presenting considerable clinical and practical value. Future research may focus on the application of semi-supervised and unsupervised learning in the field of medical imaging to enhance image utilization, improve model detection performance, and better meet clinical requirements.
1 引言
宫颈癌是一种常见的肿瘤,对女性的健康构成了严重的威胁。2020年,全球有60万宫颈癌新发病例,超过34万人死于该病[1]。宫颈癌有较长的癌前阶段,这为宫颈癌的筛查和及时治疗提供了机会。目前,检测宫颈癌最常用的方法是宫颈细胞学筛查,主要通过液基薄层细胞检测(TCT)[2]进行。具体来说,在TCT过程中,对患者的宫颈细胞进行收集并在玻片上进行染色,之后在显微镜下进行目视检查和细胞病理学分析,病理学家通过对细胞类型和形态学特征如细胞核大小、核质比等作出评估,给出初步的诊断建议。然而,这种传统的宫颈细胞学筛查费时费力且容易出错[3],同时宫颈细胞学筛查医师较为缺乏,难以满足目前宫颈癌筛查的需求[4]。随着深度学习的发展,已经有一些自动的宫颈细胞学筛查方法出现[5-6],这些方法极大地减轻了病理学家的负担,提高了检测效率。Plissiti等[7]使用VGG-16网络对单个分离的宫颈细胞图像进行分类。Talo等[8]提出了基于DenseNet161[9]的模型,对传统的方法进行了改进。Win等[10]在深度学习的基础上,引入随机森林、支持向量机等多种传统机器学习方法进行分割和分类。Du等[11]将多细胞重叠的区域划分为多个单细胞区域,并使用Faster R-CNN模型进行检测和识别。Liang等[12]提出了一个基于YOLOv3模型的全局上下文感知框架来进行异常细胞检测。为解决数据有限的问题,一种特征比较的方法[13]利用比较检测器来进行宫颈细胞检测。这些方法大多是基于卷积神经网络(CNN),由于CNN的卷积算子存在局部感受野受限的问题,其对图像中的全局特征以及远程依赖特征的提取能力不足,故这些方法的检测效果尚未达到临床要求[14]。
Transformer模型最初来源于自然语言处理领域,近年来,Transformer模型正在获得越来越多的关注。Vision Transformer模型[15]展现出Transformer模型在图像领域中的巨大潜力,其应用迅速被拓展到图像分类[16-17]、语义分割[18-19]、目标检测[20-21]等任务中。之后,又有一些新的改进的Transformer结构被提出,其中较为重要的工作是PVT模型[22]和Swin Transformer模型[16]。PVT模型引入了金字塔结构,将输入图像分解为不同分辨率图像,然后通过Transformer模型来处理这些图像,以捕获多尺度的信息。Swin Transformer模型是另一种视觉Transformer架构,通过滑动窗口操作来提高模型的局部性。
相比于自然图像,在宫颈细胞学图像中,因为异常细胞是由正常细胞缓慢发展变化而来的,异常细胞和正常细胞具有很高的相似性,所以异常细胞的检测变得非常困难。病理学家通常需要参照图像中的正常细胞,才能准确地分辨异常细胞。Transformer模型具有强大的全局特征以及远程依赖抽取能力,非常适用于宫颈异常细胞识别。然而,据我们所知,目前还只有少数的研究将Transformer模型应用到宫颈异常细胞识别上[23-24]。并且这些应用往往只是简单地用Transformer模型代替卷积架构,而没有针对特定任务进行特殊设计,因此很难达到较好的识别效果。
本文提出了一种新颖的基于Transformer模型的宫颈异常细胞检测方法,包含一种改进的Transformer编码器结构,该结构通过多尺度自注意力机制,能够更加有效地捕获图像中的关键信息,提升模型的检测效果。此外,我们也设计了一种能够自适应调整交并比(IOU)阈值的动态IOU阈值法,在训练的初始阶段,模型能够实现尽可能多的有效检测,而在训练的后期,模型能够过滤掉大部分的假阳性预测,从而提升模型的检测精度。实验表明本文的方法能够准确识别宫颈细胞学图像中的异常细胞并进行分类,模型性能优于一些现有的方法。
2 本文方法
2.1 模型整体架构
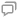
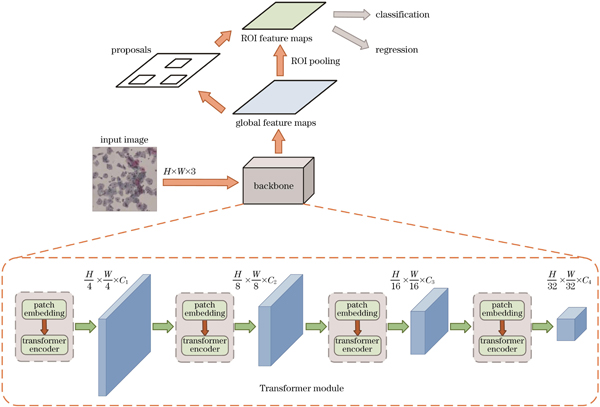
本文设计了一种能够高效进行全局特征提取的模型,整体架构如
输入图像的尺寸是H×W×3,首先将其划分为4×4×3大小的切片(patch),为了处理patch之间的位置信息,通常需要添加位置编码,以确保模型能够理解patch之间的相对位置。位置编码是一个可学习的参数矩阵,将位置信息嵌入到patch的表示中,然后将这些patch展平成线性投影,输入到第一阶段的Patch Embedding和Transformer编码器模块中,得到
由于目标检测可以分为分类任务和回归任务,因此模型使用交叉熵损失函数来处理分类任务,使用L1损失(L1 Loss,L1)函数来处理回归问题。交叉熵损失函数可以测量两个概率分布之间的差异,一般形式为
式中:
L1损失函数是一种用于回归问题的损失函数,它测量了预测值与实际值之间的绝对差异。L1损失函数的一般形式为
式中:
2.2 Transformer编码器
Transformer 编码器由多个相同结构的块(block)堆叠而成,每个block包括两部分:多头自注意力层和前馈神经网络层,每个block均执行相同的操作,具体结构如

图 2. Transformer 编码器块结构图。(a)通用结构;(b)改进结构
Fig. 2. Architecture diagrams of transformer encoder block. (a) Generic structure; (b) improved structure
Transformer模型的
式中:
每个位置的注意力分数是通过将
本文也改进了Transformer 编码器的前馈神经网络层的结构,使其更加适用于宫颈异常细胞的检测,修改后的结构如
在前馈神经网络层中也采用了Dropout处理。在每个训练迭代过程中,Dropout层随机选择一部分神经元并将其输出设置为零。这意味着在前向传播过程中,这些神经元对计算和传播梯度不起作用。Dropout层的概率参数通常表示为
2.3 动态IOU阈值
IOU阈值是指用于过滤检测框(bounding boxes)的一个阈值。IOU在目标检测中起着关键作用,主要用于衡量检测框和真实标签框之间的匹配程度,通常在0到1之间,如果IOU接近1,表示检测是准确的,如果IOU接近0,表示检测是错误的。IOU还用于目标检测的性能评估,在测试阶段可以使用IOU作为评价指标来测量检测模型的性能,例如精确率(precision)、召回率(recall)、F1分数等都依赖于IOU的计算。由此可见,IOU阈值的选择至关重要。在训练过程中,过高的IOU阈值可能会导致漏检,特别是在训练的起始阶段,低概率值的标签容易被忽略,导致模型收敛速度较慢。此外,过高的IOU阈值很容易导致小尺度目标框丢失。而过低的IOU阈值可能导致更多错误检测,因为即使检测框与真实目标的重叠很小,检测结果仍然被认为是有效的检测结果,所以假阳率增大,影响模型的检测效果。
本文提出了一种自适应的动态IOU阈值,在训练的起始阶段阈值较低,能够实现尽可能多的有效检测。而在训练的后期,较高的IOU阈值能够过滤掉大部分的假阳性预测,降低假阳率,提高模型的准确度。动态IOU阈值(
式中:
3 分析与讨论
3.1 评价指标
本文采用平均精度(AP)作为评价指标。AP是一种常用于评估目标检测和物体识别任务性能的指标,综合权衡了精确率和召回率。精确率(
式中:
AP是单个类别的精确率-召回率曲线下的面积,用于衡量模型在单个类别上的性能,范围在0到1之间。实验中也使用一些辅助性的AP指标,如AP50、AP75、APS、APM和APL。AP50是指在物体检测任务中,当IOU大于等于50%时的平均精确率。AP75是IoU大于等于75%时的平均精确率,这个指标是衡量模型在更高IoU阈值下的精确性。APS、APM、APL则是目标检测任务中小、中、大目标的AP评分。这些不同尺度的AP值可以更全面地衡量模型在不同大小物体上的性能表现。
3.2 数据集和实现细节
本文使用的全切片扫描图像(WSI)是一种高分辨率的病理组织切片的数字图像,这种图像的生成依赖于数字扫描仪和各种光学元件(如光学镜头、光学传感器和透镜)。将光聚焦在组织切片上,利用各种光学器件将玻片上的光学信息转为数字图像,保留了病理学上的详细结构,使医生能够直接观察组织和细胞的形态、结构和染色情况。
在20倍的物镜放大倍数下,通过明场扫描的方法对苏木精-伊红(H&E)染色的切片进行扫描,所获得的全切片图像是一种多分辨率的层次模型,其包含更多的信息,这是传统的光学显微镜无法实现的。
本文使用的数据集来自天池宫颈癌风险智能诊断挑战赛[26]。所有图像均为宫颈癌液基薄层细胞,所有标签均由专业医师标注[27]。该数据集包含800张WSI,其中500张为包含异常细胞的阳性WSI,我们随机选择其中的350张用来训练,75张作为验证集,剩下的作为测试集。由于WSI尺寸极大,不能直接输入到模型中,首先需要对其中包含异常细胞的局部图像进行裁剪,裁剪尺寸固定为1000 pixel
由于医学领域的有标签数据相对稀缺,为了最大程度利用现有的有标签数据,在训练之前需要对输入图像进行充分的数据增强。本文主要采用两个类型的增强方法,包括形状变换和颜色变换。对于形状变换,采用旋转、镜像翻转、平移和缩放变换等操作,对输入图像的形状进行调整,增加模型对不同视角和方向图像的识别能力。对于颜色变换,主要采用亮度和对比度调整、模糊及滤波等操作,在保持数据真实性和有效性的前提下,对图像进行尽可能多的变换。这些数据增强方法在医学图像领域中尤为重要,因为医学图像往往受多种因素的影响,如设备差异、光照变化等,所以医学图像的风格差异较大。通过应用数据增强技术,可以增加医学图像数据的多样性,提高深度学习模型的鲁棒性和泛化性能。
本文代码基于PyTorch框架,使用8个NVIDIA GeForce RTX 2080 Ti显卡进行训练。优化器选择了随机梯度下降(SGD)算法,并设置动量参数为0.9,初始学习率为0.01。
3.3 对比实验
为了验证本文方法的有效性,我们在所使用的天池数据集上开展了实验,并与常见的通用目标检测方法进行了对比。为了保证对比实验的公平性,所有模型都在相同的环境中进行训练和测试,实验结果如
表 1. 各种模型的实验结果
Table 1. Experimental results of various models
|
除了与通用模型进行对比外,本文模型还与专门为宫颈异常细胞检测而设计的网络attFPN[35]进行了比较。attFPN由两个部分组成:一是模拟病理学家阅读宫颈细胞学图像的注意力模块,能够对提取到的特征进行细化以强调或抑制某些特征;另一个模块是多尺度特征融合网络,通过融合细化后的特征,检测不同区域的宫颈癌变细胞。其检测结果如
表 2. 本文模型和attFPN的对比
Table 2. Comparison between proposed model and attFPN
|
本文方法和其他方法的可视化结果对比如

图 5. 不同模型的检测效果对比。(a)真实标签;(b)所提方法;(c)Sparse R-CNN
Fig. 5. Comparison of detection effects of different models. (a) Ground truth; (b) proposed method; (c) Sparse R-CNN
3.4 消融实验
消融实验主要是为了验证Transformer作为backbone时的效果,以及我们改进的Transformer 编码器模块和动态IOU阈值的有效性。
表 3. 不同backbone选择下的实验结果对比
Table 3. Comparison of experimental results under different backbone choices
|
针对改进的Transformer 编码器模块和动态IOU阈值的消融实验结果如
表 4. 不同模块的消融实验结果
Table 4. Ablation experiment results of different modules
|
我们也对动态IOU阈值和多个固定的IOU阈值进行了定量对比分析,固定的IOU阈值分别取0.5、0.6、0.7,结果如
表 5. 动态IOU阈值和多个固定IOU阈值模型的对比
Table 5. Comparison among models based on dynamic IOU threshold and multiple fixed IOU thresholds
|
此外,我们也比较了动态IOU阈值和固定IOU阈值模型在训练过程中的损失和AP的变化,如

图 6. 动态IOU阈值和固定IOU阈值下的损失和AP。(a)损失;(b)AP
Fig. 6. Loss and AP under dynamic IOU threshold and fixed IOU threshold. (a) Loss; (b) AP
除了定量的比较外,也对模型的有效性进行了定性分析。本文采用Grad-CAM[36]生成热图,以此对模型的检测能力进行比对。热图可以提供关于目标位置的精细粒度信息,使模型可以更准确地理解图像中的目标分布情况。将不采用改进Transformer编码器和动态IOU阈值的原始Transformer模型和本文模型进行对比,生成的热图如

图 7. 消融实验热图。(a)原始图像;(b)原始Transformer模型生成的热图;(c)我们的方法生成的热图
Fig. 7. Ablation experiment heatmaps. (a) Original images; (b) heatmap generated by original Transformer model; (c) heatmap generated by our method
4 结论
提出了一种宫颈异常细胞自动识别方法,模型基于Transformer架构。在此基础上提出了改进的Transformer 编码器结构和可以动态变化的IOU阈值。在数据集上进行了多种对比实验,结果表明:所提出的方法在精度等各种指标上均优于已有的方法,能够实现准确的宫颈异常细胞识别。通过消融实验,证明了所提出的两个模块均能够提高模型对宫颈异常细胞的识别准确度。总的来看,所提出的方法能够极大地提升医学图像的筛查效率,节省医疗时间和资源,及时发现癌症病变,具有一定的临床和使用价值。
后续的研究将更关注数据的高效利用。医学图像领域的有标注数据集较为稀缺,这极大地限制了模型性能的提升。然而在医学领域中还存在着大量的无标签或弱标签数据,这些数据也蕴含着相当多的可以利用的特征,因此无标签数据的利用对于医学图像领域的模型改进十分必要。后续的研究可以更多关注半监督学习和无监督学习在医学图像领域中的应用,利用这些方法提高医学图像的利用率,提升模型的检测效果,使其更好地符合临床需求。
[1] Sung H, Ferlay J, Siegel R L, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA: A Cancer Journal for Clinicians, 2021, 71(3): 209-249.
[2] de Bekker-Grob E W, de Kok I M C M, Bulten J, et al. Liquid-based cervical cytology using ThinPrep technology: weighing the pros and cons in a cost-effectiveness analysis[J]. Cancer Causes & Control, 2012, 23(8): 1323-1331.
[3] Elsheikh T M, Austin R M, Chhieng D F, et al. American Society of Cytopathology workload recommendations for automated Pap test screening: developed by the productivity and quality assurance in the era of automated screening task force[J]. Diagnostic Cytopathology, 2013, 41(2): 174-178.
[4] 李雪, 石中月, 杨志明, 等. 人工智能辅助分析在宫颈液基薄层细胞学检查中的应用价值[J]. 首都医科大学学报, 2020, 41(3): 360-363.
Li X, Shi Z Y, Yang Z M, et al. Value about artificial intelligence-assisted liquid-based thin-layer cytology for cytology cervical cancer screening[J]. Journal of Capital Medical University, 2020, 41(3): 360-363.
[5] Chen Y F, Huang P C, Lin K C, et al. Semi-automatic segmentation and classification of pap smear cells[J]. IEEE Journal of Biomedical and Health Informatics, 2014, 18(1): 94-108.
[6] William W, Ware A, Basaza-Ejiri A H, et al. A review of image analysis and machine learning techniques for automated cervical cancer screening from pap-smear images[J]. Computer Methods and Programs in Biomedicine, 2018, 164: 15-22.
[7] PlissitiM E, DimitrakopoulosP, SfikasG, et al. Sipakmed: a new dataset for feature and image based classification of normal and pathological cervical cells in pap smear images[C]∥2018 25th IEEE International Conference on Image Processing (ICIP), October 7-10, 2018, Athens, Greece. New York: IEEE Press, 2018: 3144-3148.
[8] Talo M. Diagnostic classification of cervical cell images from pap smear slides[J]. Academic Perspective Procedia, 2019, 2(3): 1043-1050.
[9] HuangG, LiuZ, Van Der MaatenL, et al. Densely connected convolutional networks[C]∥2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), July 21-26, 2017, Honolulu, HI, USA. New York: IEEE Press, 2017: 2261-2269.
[10] Win K P, Kitjaidure Y, Hamamoto K, et al. Computer-assisted screening for cervical cancer using digital image processing of pap smear images[J]. Applied Sciences, 2020, 10(5): 1800.
[11] Du, LiX Y, LiQ H. Detection and classification of cervical exfoliated cells based on faster R-CNN[C]∥2019 IEEE 11th International Conference on Advanced Infocomm Technology (ICAIT), October 18-20, 2019, Jinan, China. New York: IEEE Press, 2019: 52-57.
[12] Liang Y X, Pan C L, Sun W X, et al. Global context-aware cervical cell detection with soft scale anchor matching[J]. Computer Methods and Programs in Biomedicine, 2021, 204: 106061.
[13] Liang Y X, Tang Z H, Yan M, et al. Comparison detector for cervical cell/clumps detection in the limited data scenario[J]. Neurocomputing, 2021, 437: 195-205.
[14] 辛仲宏, 雷军强, 郭城, 等. 深度学习用于宫颈癌诊疗研究进展[J]. 中国医学影像技术, 2022, 38(5): 779-782.
Xin Z H, Lei J Q, Guo C, et al. Research progresses of deep learning in diagnosis and treatment of cervical cancer[J]. Chinese Journal of Medical Imaging Technology, 2022, 38(5): 779-782.
[16] LiuZ, LinY T, CaoY, et al. Swin transformer: hierarchical vision transformer using shifted windows[C]∥2021 IEEE/CVF International Conference on Computer Vision (ICCV), October 10-17, 2021, Montreal, QC, Canada. New York: IEEE Press, 2022: 9992-10002.
[17] PanX R, GeC J, LuR, et al. On the integration of self-attention and convolution[C]∥2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 18-24, 2022, New Orleans, LA, USA. New York: IEEE Press, 2022: 805-815.
[18] StrudelR, GarciaR, LaptevI, et al. Segmenter: transformer for semantic segmentation[C]∥2021 IEEE/CVF International Conference on Computer Vision (ICCV), October 10-17, 2021, Montreal, QC, Canada. New York: IEEE Press, 2022: 7242-7252.
[19] HadsellR, ChopraS, LeCunY. Dimensionality reduction by learning an invariant mapping[C]∥2006 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’06), June 17-22, 2006, New York, NY, USA. New York: IEEE Press, 2006: 1735-1742.
[20] LiY H, MaoH Z, GirshickR, et al. Exploring plain vision transformer backbones for object detection[M]∥AvidanS, BrostowG, CisséM, et al. Computer vision-ECCV 2022. Lecture notes in computer science. Cham: Springer, 2022, 13669: 280-296.
[21] PanX R, XiaZ F, SongS J, et al. 3D object detection with pointformer[C]∥2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 20-25, 2021, Nashville, TN, USA. New York: IEEE Press, 2021: 7459-7468.
[22] WangW H, XieE Z, LiX, et al. Pyramid vision transformer: a versatile backbone for dense prediction without convolutions[C]∥2021 IEEE/CVF International Conference on Computer Vision (ICCV), October 10-17, 2021, Montreal, QC, Canada. New York: IEEE Press, 2022: 548-558.
[23] Liu W L, Li C, Xu N, et al. CVM-Cervix: a hybrid cervical Pap-smear image classification framework using CNN, visual transformer and multilayer perceptron[J]. Pattern Recognition, 2022, 130: 108829.
[24] Liu Y T, Zhao J J, Luo Q Y, et al. Automated classification of cervical lymph-node-level from ultrasound using Depthwise Separable Convolutional Swin Transformer[J]. Computers in Biology and Medicine, 2022, 148: 105821.
[25] GirshickR. Fast R-CNN[C]∥2015 IEEE International Conference on Computer Vision (ICCV), December 7-13, 2015, Santiago, Chile. New York: IEEE Press, 2016: 1440-1448.
[26] ‘Dataset’[EB/OL]. [2023-05-06]. https:∥tianchi.aliyun.com/competition/.
[27] 梁義钦, 赵司琦, 王海涛, 等. 两阶段分析的异常簇团宫颈细胞检测方法[J]. 哈尔滨理工大学学报, 2022, 27(2): 76-84.
Liang Y Q, Zhao S Q, Wang H T, et al. Two-stage detection method for abnormal cluster cervical cells[J]. Journal of Harbin University of Science and Technology, 2022, 27(2): 76-84.
[28] LinT Y, GoyalP, GirshickR, et al. Focal loss for dense object detection[C]∥2017 IEEE International Conference on Computer Vision (ICCV), October 22-29, 2017, Venice, Italy. New York: IEEE Press, 2017: 2999-3007.
[29] CarionN, MassaF, SynnaeveG, et al. End-to-end object detection with transformers[M]∥VedaldiA, BischofH, BroxT, et al. Computer vision–ECCV 2020. Lecture notes in computer science. Cham: Springer, 2020, 12346: 213-229.
[30] TianZ, ShenC H, ChenH, et al. FCOS: fully convolutional one-stage object detection[C]∥2019 IEEE/CVF International Conference on Computer Vision (ICCV), October 27-November 2, 2019, Seoul, Korea (South). New York: IEEE Press, 2020: 9626-9635.
[32] SunP Z, ZhangR F, JiangY, et al. Sparse R-CNN: end-to-end object detection with learnable proposals[C]∥2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 20-25, 2021, Nashville, TN, USA. New York: IEEE Press, 2021: 14449-14458.
[33] CaiZ W, VasconcelosN. Cascade R-CNN: delving into high quality object detection[C]∥2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, June 18-23, 2018, Salt Lake City, UT, USA. New York: IEEE Press, 2018: 6154-6162.
[34] ZhengA L, ZhangY A, ZhangX Y, et al. Progressive end-to-end object detection in crowded scenes[C]∥2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 18-24, 2022, New Orleans, LA, USA. New York: IEEE Press, 2022: 847-856.
[35] Cao L, Yang J Y, Rong Z W, et al. A novel attention-guided convolutional network for the detection of abnormal cervical cells in cervical cancer screening[J]. Medical Image Analysis, 2021, 73: 102197.
[36] SelvarajuR R, CogswellM, DasA, et al. Grad-CAM: visual explanations from deep networks via gradient-based localization[C]∥2017 IEEE International Conference on Computer Vision (ICCV), October 22-29, 2017, Venice, Italy. New York: IEEE Press, 2017: 618-626.
Article Outline
张峥, 陈明销, 李新宇, 程逸, 申书伟, 姚鹏. 基于Transformer的宫颈异常细胞自动识别方法[J]. 中国激光, 2024, 51(3): 0307108. Zheng Zhang, Mingxiao Chen, Xinyu Li, Yi Chen, Shuwei Shen, Peng Yao. Automatic Identification of Cervical Abnormal Cells Based on Transformer[J]. Chinese Journal of Lasers, 2024, 51(3): 0307108.