自适应混合发射单分子定位算法
Currently, various super-resolution imaging technologies can surpass the Abbe diffraction limit, thereby improving imaging resolution to several tens of nanometers. This provides biologists with an effective tool for investigating biological structures and their functions on a novel scale. Among these, single-molecule localization techniques such as photoactivated localization microscopy (PALM) and stochastic optical reconstruction microscopy (STORM) yield the highest resolution. Traditional fitting-based methods, such as single-emitter localization (SE) and multi-emitter localization (ME) algorithms, employ fixed-size sliding windows to select the fitting areas. However, this was found to lead to an inadequate use of the prior emitter recognition information during the emitter localization stage in this study, thereby resulting in diverse advantageous density ranges and different artifact forms of SE and SM. The SE results are distorted by truncates near the emitters, which are generated by the fixed sizes of the fitting areas, whereas the ME suffers from an inappropriate fitting number. In summary, a self-adaptive mixed-emitter single-molecule localization algorithm (SM) that can adaptively determine the fitting area and fitting number is proposed in this study. Consequently, compared with the SE and ME algorithms, the images reconstructed by the SM algorithm exhibit a superior resolution and contrast over the complete density range on both simulated and experimental data.
The complete SM algorithm comprises several steps. First, an SNR binary map that can shrink and expand with the power of noise was generated based on the original image. Subsequently, the SNR binary map was combined with the local maxima for emitter recognition, and the sliding window and fitting number were generated using the SNR binary map. The center and size of the generated sliding window were then determined based on the center position and size of the connected domain, respectively, whereas the fitting number was obtained from previous emitter recognition results. Subsequently, maximum likelihood estimation (MLE) or least squares (LS) fitting was performed in each fitting area to obtain the subpixel positions. Finally, the performance of the SM algorithm was investigated using simulated and experimental data.
Under a low or high labeling density, the SM algorithm can effectively reduce crosstalk and mismatch errors, which promotes the recovery of super-resolution images closer to the synthesized benchmark images compared to those recovered by the SE and ME algorithms (Fig. 1). For a low labeling density, the SM algorithm exhibits a slightly better precision, recall, Jaccard index, and RMSE than the SE algorithm, and significantly superior results compared to those of the ME algorithm. With an increasing labeling density, the SM algorithm is marginally inferior to the ME algorithm in terms of the precision, recall, and Jaccard index, but is still significantly better than those of the SE algorithm. In terms of the RMSE, the SM and SE algorithms exhibit comparable localization errors, which are both worse than those of the ME algorithm [Figs. 3(a)?(c)]. Quantitative comparisons between the synthesized benchmark images and super-resolution images recovered by the different algorithms are performed using three indicators: PSNR, SSIM, and RMSE. The SM algorithm produces images with a higher similarity to the ground truth, as indicated by all three indicators (Table 1). In addition, it also successfully restores the structure with an interval of 20 nm, which is not achieved using the SE and ME algorithms [Figs. 3(d)?(e)]. On the α-tubulin dataset labeled as Alexa Fluor 647, the SM algorithm outperforms both the SE and ME algorithms in terms of resolution and contrast, as calculated using the FRC metrics (Fig. 4 and Table 2).
In this study, a self-adaptive mixed-emitter single-molecule localization algorithm that enables the adaptive determination of the fitting area and fitting number is proposed. Compared to the SE and ME algorithms, the SM algorithm can significantly reduce the artifacts caused by mismatch and crosstalk errors, resulting in an enhanced resolution and contrast within the full applicable density range of the fitting method. In terms of the speed, the current SM algorithm is faster than ME algorithm by a factor of 3?4, and slower than the SE algorithm by one order of magnitude. However, the number of fitting iterations required by the SM algorithm is the same as that required by the SE algorithm. Therefore, after optimization, the SM algorithm has the potential to achieve a speed comparable to that of the SE algorithm. Although the analysis and experiments in this study were conducted under two-dimensional and single-channel conditions, the inherent mechanism of the SM method allows for its easy integration with more complex single-molecule imaging technologies, such as three-dimensional and multi-channel situations. In future research, the SM algorithm should be further refined and its reliability and stability should be verified, thereby expanding its advantages in the field of biological imaging.
1 引言
超分辨荧光显微镜可以突破阿贝衍射极限的限制,将分辨率提升至几十纳米,为生物学家从更微观的尺度研究生物结构和功能提供了强有力的帮助[1-4]。单分子定位显微镜是其中最具代表性的显微镜[5-8],其原理是利用开关荧光蛋白在开态和关态间的切换[9],在单次曝光时间内随机激发少部分荧光蛋白,获得稀疏的荧光图像,随后将这一过程重复上万次,从而获得足够的荧光标记密度。单分子定位成像的质量与荧光标记密度、定位精度有关[10-11],因而荧光分子利用效率和定位精度是单分子定位成像质量的重要指标。目前,针对精度和速度的提高已经有相当数量的研究,并取得了丰富的研究成果[12-19]。总体而言,基于拟合的方法可以分为单发射(SE)拟合和多发射(ME)拟合算法。其中:SE算法假设每个拟合区域内只存在一个需要拟合的荧光点,模型简单且定位速度快,但会拒绝不满足单发射的情况,因而荧光分子利用率低;ME算法假设每个拟合区域存在多个需要拟合的荧光点,通过计算得到应拟合点的数目,模型约束更少、失效风险小,因而荧光分子利用率高。在实际应用中,SE拟合和ME拟合总是表现出不同的优势密度范围。笔者针对这一现象进行了研究,并提出了串扰误差与失配误差的概念。
由于样品荧光标记密度的空间分布总是非均匀的,受串扰误差和失配误差的影响,全局使用SE、ME算法难以达到全密度范围内的最优定位效果,且会不可避免地带来光子利用效率低下和计算速度慢的问题。因此,笔者提出了自适应混合发射单分子定位(SM)算法,创新性地采用自适应滑窗而非传统的固定大小滑窗进行拟合区域选取,并根据拟合区域内识别的荧光分子数选择定位模型,从而避免了上述两种误差的产生。验证结果显示:SM算法在所有密度情况下恢复的图像分辨率和对比度均优于SE和ME算法,且速度比ME算法快3~4倍。
2 基本原理
2.1 串扰误差和失配误差
通过对SE及ME算法的定位误差进行研究,笔者发现其主要包括串扰误差和失配误差。前者是因为两种算法都采用固定大小的滑窗进行拟合区域的选取,因此定位结果容易受到相邻荧光点的干扰。串扰误差主要发生在高密度情况下,同时在ME拟合中表现得相对较轻,因为ME算法可以将串扰荧光点进行定位从而排除其影响。后者是因为其所需拟合的荧光分子数目未知,易出现实际拟合数目和发射数目不匹配的情况。值得注意的是,失配误差发生在ME拟合的全密度范围内。在低发射密度情况下,SE拟合几乎不受串扰误差的影响,而ME拟合则受失配误差的影响,因而SE拟合的定位精度高;在高发射密度情况下,SE拟合会受到串扰误差的严重影响,而ME拟合受失配误差影响的程度相对较轻,因此ME拟合的定位精度高。
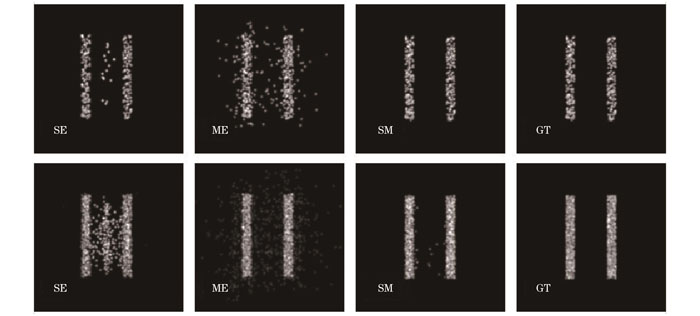

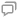
图 1. 不同算法对标记密度为1
Fig. 1. Reconstructed images of two 200 nm stripes apart generated by different algorithms, density of the strip is 1
2.2 单分子定位成像模型和信噪比二值图
单分子定位显微镜采集到的原始图像
式中:
信噪比二值图生成步骤如下:首先用分位数为
信噪比二值图同像素附近的整体信噪比密切相关。假设第一步二值化灰度为0的像素全部来源于噪声点,则对于给定的
式中:
在荧光信号强度为
若定义
设
对于给定的邻域
式中:m表示生成信噪比二值图过程中的强度分位数。
2.3 荧光点识别算法及自适应滑窗定位
荧光点识别方法通常都是将局部极值与阈值条件相结合,区别仅在于预处理及确定阈值的方法不同[20-22]。但阈值条件只从孤立的像素强度出发,且在找寻局部极值前需要对图像进行平滑滤波处理,导致难以识别部分重叠的荧光点。SM算法利用“信噪比二值图”反映出的空间连续性特征进行荧光点的识别,同时位于原始图像局部极值处和信噪比二值图中“1”处的像素被识别为荧光点。由于高信噪比处的图像基本由荧光信号决定,所以SM算法不需要进行平滑滤波来抑制噪声,可以直接在原始图像上找寻局部极值,因此对重叠荧光点的分辨能力强。算法流程如
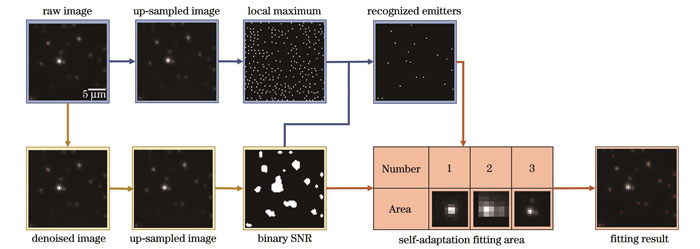
图 2. SM算法流程图,其中黄色、蓝色和橙色箭头分别展示了生成信噪比二值图、荧光点识别、自适应拟合算法的过程
Fig. 2. Flowchart of SM algorithm, where the process of binary SNR generation, emitter identification and self-adaptation fitting algorithm are represented by yellow, blue and orange arrows, respectively
在荧光点定位方面,与采用固定大小滑窗进行定位的SE算法和ME算法不同,SM算法采用自适应滑窗,流程如
3 分析与讨论
3.1 仿真数据上的算法对比
采用召回率、准确率、Jaccard系数、均方根误差(RMSE)[23]作为仿真实验的评价指标。前三个指标从集合角度进行考察:召回率表示检出真实荧光点数目与全体荧光点数目之比,准确率表示检出真实荧光点数目与检出荧光点数目之比,Jaccard系数表示检出荧光点与真实荧光点交集同检出荧光点与真实荧光点并集之比。均方根误差从距离角度对定位精度进行考察。仿真数据包括信噪比为10、密度在0.1~2
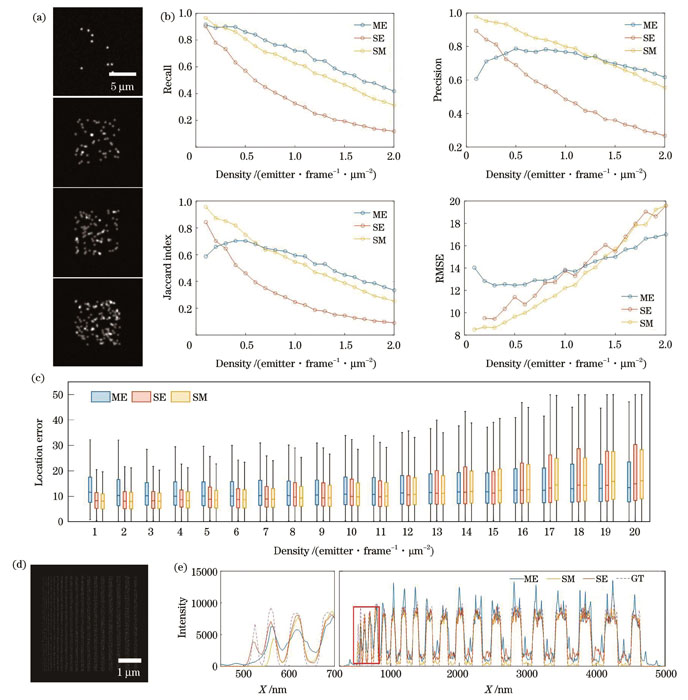
图 3. 仿真数据及仿真数据上的算法对比。(a)生成的不同密度的仿真图像;(b)不同算法的指标比较;(c)不同算法的定位误差;(d)仿真的参考图像;(e)不同算法生成的超分辨图像的横截面强度及其局部放大图
Fig. 3. Simulation data and algorithm comparison on simulation data. (a) Simulated images with different densities; (b) comparison of metrics of different algorithms; (c) localization error of different algorithms; (d) simulated reference image (ground truth,GT); (e) sectional intensity profiles and local magnification of reconstructed super-resolution images by different algorithms
由
为了更直观地比较不同算法重构出的超分辨图像的质量,通过仿真生成了
表 1. 不同算法恢复的超分辨图像与参考图像的对比
Table 1. Comparison of super-resolution images recovered from different algorithms with ground truth image
|
3.2 实验数据上的算法对比
在Alexa Fluor 647标记的α微管蛋白数据上进行算法的比较,实验激发光功率为2
在分辨率方面,利用滚动傅里叶环相关(rFRC)算法[24]得到不同算法恢复的超分辨图像的局部分辨率(如
表 2. rFRC给出的三种算法分辨率的定量结果
Table 2. Quantitative resolution features of different algorithms given by rFRC
|
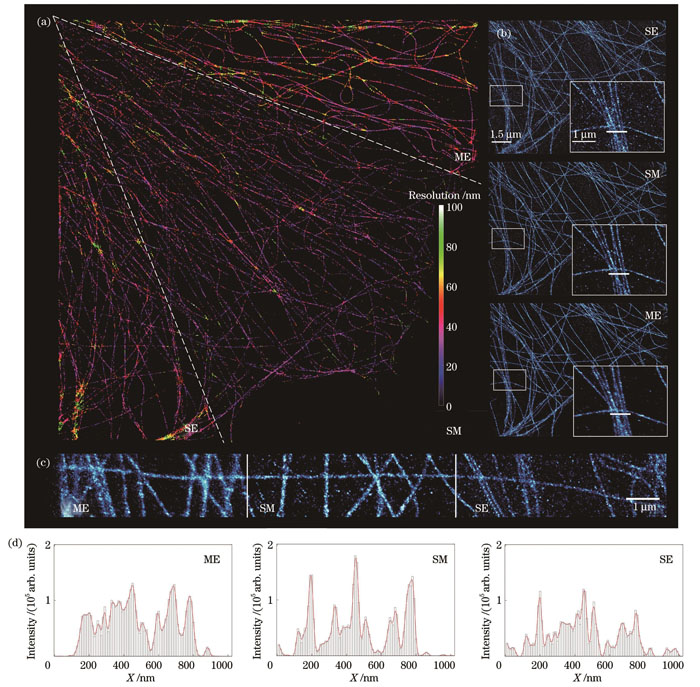
图 4. 实验数据上的算法对比。(a)不同算法获得的超分辨图像的rFRC图;(b)~(c)局部放大视图;(d)图4(b)中白线标注部位横截面强度的对比
Fig. 4. Algorithm comparison on experiment data. (a) rFRC maps of super-resolution images obtained from different algorithms; (b)‒(c) local magnification of super resolution images; (d) sectional intensity profile corresponding to white lines in Fig. 4(b)
3.3 算法特点对比
通过对除随机误差以外的其余可能误差来源进行理论分析以及实验对比可以发现,本文所提SM算法具有更少的误差来源、全密度范围内的优秀定位结果、较高的光子利用效率。SM算法与SE、ME算法的对比如
表 3. SE、ME、SM算法的对比
Table 3. Comparison of SE, ME and SM algorithms
|
4 结论
本文提出了自适应混合发射单分子定位算法,该算法可自适应调节拟合区域的大小并可在拟合区域内选择合适的模型初值。在分辨率方面,SM算法能够避免由固定滑窗引起的串扰误差以及ME算法导致的失配误差,在拟合方法适用的全密度范围内表现出了更高的分辨率和对比度,弥补了SE算法和ME算法分别只在部分密度范围具有定位优势的不足。在速度方面,SM算法比ME算法快3~4倍,比SE算法慢一个数量级。理论上,该算法所需的拟合次数同SE算法相同,比ME算法少一个数量级。因此,该算法经过优化后有可能达到同SE算法相近的速度。
综上所述,为了更好地满足生物学家对成像实验的需求,单分子定位技术应在成像速度、精度和稳定性等方面持续优化,包括在成像端进行改进[25]。同时,也应在三维、多色和活细胞成像方向不断发展[26-27]。所提算法目前应用在二维和单通道情况下,但是从内在机制上看,该算法同样可以与三维、多通道等更复杂的单分子成像技术相结合。笔者拟继续改进该算法模型,提升其可靠性和稳定性,使其发挥出更大作用。
[1] Hell S W. Far-field optical nanoscopy[J]. Science, 2007, 316(5828): 1153-1158.
[2] Huang B, Babcock H, Zhuang X W. Breaking the diffraction barrier: super-resolution imaging of cells[J]. Cell, 2010, 143(7): 1047-1058.
[3] Zeng Z P, Xie H, Chen L, et al. Computational methods in super-resolution microscopy[J]. Frontiers of Information Technology & Electronic Engineering, 2017, 18(9): 1222-1235.
[4] 高露, 高贝贝, 王富. 超分辨显微成像技术在活体大脑成像中的应用[J]. 中国激光, 2022, 49(20): 2007301.
[5] Betzig E, Patterson G H, Sougrat R, et al. Imaging intracellular fluorescent proteins at nanometer resolution[J]. Science, 2006, 313(5793): 1642-1645.
[6] Hess S T, Girirajan T P K, Mason M D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy[J]. Biophysical Journal, 2006, 91(11): 4258-4272.
[7] Rust M J, Bates M, Zhuang X W. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM)[J]. Nature Methods, 2006, 3(10): 793-796.
[8] 李雨竹, 李传康, 郝翔, 等. 基于单分子定位成像技术的研究及展望[J]. 激光与光电子学进展, 2020, 57(24): 240002.
[9] Zhang M S, Fu Z F, Li C Q, et al. Fast super-resolution imaging technique and immediate early nanostructure capturing by a photoconvertible fluorescent protein[J]. Nano Letters, 2020, 20(4): 2197-2208.
[10] van de Linde S, Wolter S, Heilemann M, et al. The effect of photoswitching kinetics and labeling densities on super-resolution fluorescence imaging[J]. Journal of Biotechnology, 2010, 149(4): 260-266.
[11] Nieuwenhuizen R P J, Lidke K A, Bates M, et al. Measuring image resolution in optical nanoscopy[J]. Nature Methods, 2013, 10(6): 557-562.
[12] Small A, Stahlheber S. Fluorophore localization algorithms for super-resolution microscopy[J]. Nature Methods, 2014, 11(3): 267-279.
[13] Cheezum M K, Walker W F, Guilford W H. Quantitative comparison of algorithms for tracking single fluorescent particles[J]. Biophysical Journal, 2001, 81(4): 2378-2388.
[14] Parthasarathy R. Rapid, accurate particle tracking by calculation of radial symmetry centers[J]. Nature Methods, 2012, 9(7): 724-726.
[15] Thompson R E, Larson D R, Webb W W. Precise nanometer localization analysis for individual fluorescent probes[J]. Biophysical Journal, 2002, 82(5): 2775-2783.
[16] Smith C S, Joseph N, Rieger B, et al. Fast, single-molecule localization that achieves theoretically minimum uncertainty[J]. Nature Methods, 2010, 7(5): 373-375.
[17] 杨建宇, 董浩, 邢福临, 等. 单分子定位超分辨成像技术进展及应用[J]. 激光与光电子学进展, 2021, 58(12): 1200001.
[18] Li L C, Xin B, Kuang W B, et al. Divide and conquer: real-time maximum likelihood fitting of multiple emitters for super-resolution localization microscopy[J]. Optics Express, 2019, 27(15): 21029-21049.
[19] Huang F, Schwartz S L, Byars J M, et al. Simultaneous multiple-emitter fitting for single molecule super-resolution imaging[J]. Biomedical Optics Express, 2011, 2(5): 1377-1393.
[20] Holden S J, Uphoff S, Kapanidis A N. DAOSTORM: an algorithm for high- density super-resolution microscopy[J]. Nature Methods, 2011, 8(4): 279-280.
[21] Ovesný M, Křížek P, Borkovec J, et al. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging[J]. Bioinformatics, 2014, 30(16): 2389-2390.
[22] Li Y M, Ishitsuka Y, Hedde P N, et al. Fast and efficient molecule detection in localization-based super-resolution microscopy by parallel adaptive histogram equalization[J]. ACS Nano, 2013, 7(6): 5207-5214.
[23] Sage D, Pham T A, Babcock H, et al. Super-resolution fight club: assessment of 2D and 3D single-molecule localization microscopy software[J]. Nature Methods, 2019, 16(5): 387-395.
[25] Gu L S, Li Y Y, Zhang S W, et al. Molecular-scale axial localization by repetitive optical selective exposure[J]. Nature Methods, 2021, 18(4): 369-373.
[26] Li Y M, Mund M, Hoess P, et al. Real-time 3D single-molecule localization using experimental point spread functions[J]. Nature Methods, 2018, 15(5): 367-369.
[27] Zhang Y D, Schroeder L K, Lessard M D, et al. Nanoscale subcellular architecture revealed by multicolor three-dimensional salvaged fluorescence imaging[J]. Nature Methods, 2020, 17(2): 225-231.
Article Outline
刘一哲, 赵唯淞, 刘宇桢, 李浩宇. 自适应混合发射单分子定位算法[J]. 中国激光, 2023, 50(21): 2107106. Yizhe Liu, Weisong Zhao, Yuzhen Liu, Haoyu Li. Self-Adaptive Mixed-Emitter Single-Molecule Localization Algorithm[J]. Chinese Journal of Lasers, 2023, 50(21): 2107106.






